|
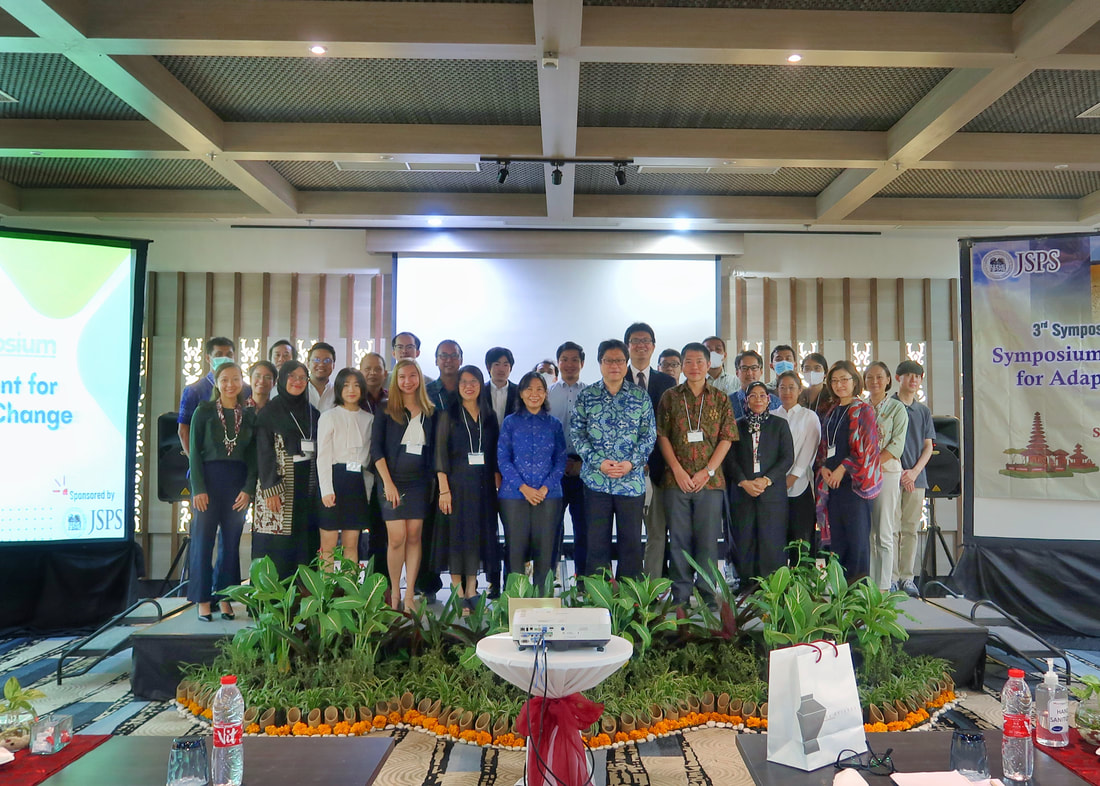
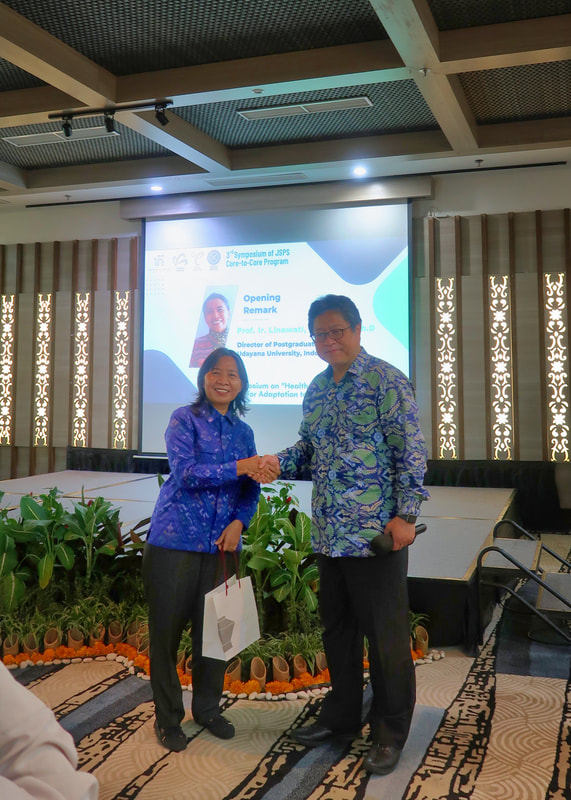
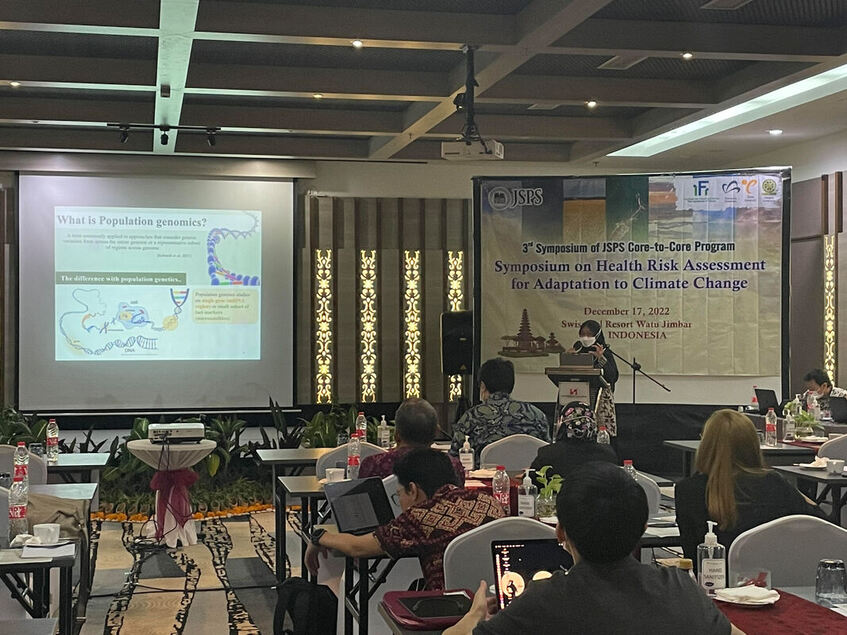
The 3rd Symposium of the JSPS Core-to-Core program was conducted at Swiss-Bel Resort, Bali, Indonesia last December 17, 2022. The symposium was organized by The University of Tokyo (IFI), Yamagata University, Ehime University, and Udayana University. The goal of the symposium was to discuss how climate change impacts key areas i.e. health, infectious disease prevention, nutrition, food security, and sustainability. Prof. Kensuke Fukushi (The University of Tokyo, IFI) and Prof. lr. Linawati (Udayana University) delivered the welcome remarks followed by four main sessions: (1) climate change impact moderated by Prof. Kensuke Fukushi, (2) environment moderated by Prof. Toru Watanabe, (3) health, nutrition, and food moderated by Prof. Kozo Watanabe, and (4) early career and student: climate, environment & health, nutrition & food moderated by Tri Atmaja (Ph.D. student at the University of Tokyo). The first three sessions of the symposium tackled the global issues concerning food production, food security, water quality and access, agriculture, mosquito vectors, and mosquito-borne diseases. These sessions highlighted the need to accelerate efforts that can help minimize the risks and threats of long-term shifts in temperatures and weather patterns. Both mitigation and adaptation should be considered when developing new initiatives to resolve these issues. The final session focused on early career scientists and students who were given the opportunity to share their researches. One research aimed to use a remote sensing analysis for the assessment of coastal flood loss and damage linked to climate change whereas other researches were geared towards strengthening food system resilience, sustainable food consumption, and disease prevention. Atikah Fitria Muharommah and Jerica Reyes, Ph.D. students from MECOH lab, presented their works on the population genetic structure of Aedes aegypti as well as the use of Wolbachia and CRISPR-Cas 9 tool for understanding Dengue infection within the vector host. The symposium served as an avenue for the scientific discussion of problems caused by climate change. Also, It was a way to establish collaboration between attending institutions and mobilize solutions for the attainment of a climate neutral environment. Credits/Disclaimer: Some photos were obtained from the organizing committee and were only used for the purpose of documentation.
0 Comments
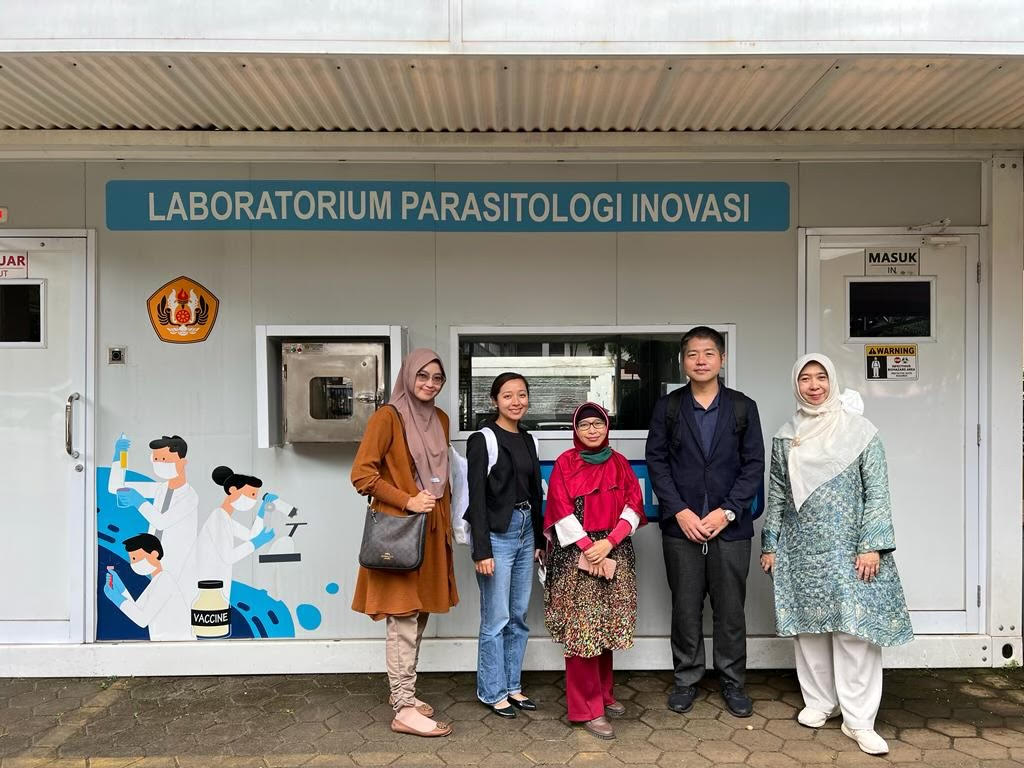
Dr. Kozo Watanabe and Jerica Reyes (Ph.D. student) visited the Universitas Padjadjaran in Bandung Indonesia to discuss their long-standing collaboration. For the past years, the MECOH lab has conducted projects focusing on arthropod-borne diseases together with Dr. Lia Faridah, Dr. Savira Ekawardhani, and Dr. Nisa Fauziah. They discussed future research collaboration in the meeting and the prospect of establishing a satellite laboratory under Ehime University at the Universitas Padjadjaran. Dr. Ruswana Anwar, Vice Dean for Academic, Student Affair, and Research in the Universitas Padjadjaran attended the meeting to exchange ideas regarding the satellite laboratory. One of the main goals of the satellite laboratory is to strengthen the international collaborative research between Ehime University and Universitas Padjadjaran. Both universities intend to collaborate on researches involving Dengue, Malaria, and Anti-microbial Resistance (AMR). The two universities plan to merge their research goals to resolve global issues on both health and the environment. The meeting also included a tour of the research facilities in the Universitas Padjadjaran. Their facilities include a cell culture room, cytogenetics laboratory, immunology laboratory, next-generation sequencing, and a MoVi Labtainer which is a recent addition to handle COVID-19 samples. To conclude the day, everyone enjoyed some of the local Indonesian food!
Four students from two universities in Indonesia spent a month in the MECOH lab as interns. Mia and Naomi from Institut Teknologi Bandung and Sidah and Vita from Universitas Gadjah Mada Indonesia worked with some of the Ph.D. students in our lab to learn different molecular techniques. Here, they talked about their lab experience and learnings: Mia: Thank you MECOH Lab, I learned a lot about the environment from the scientist's point of view, supported by complete lab equipment. This will be useful for me in the future, especially for the sanitation sector which I have been studying on so far. Thanks to sensei and the students for all the knowledge. In addition, many foreign students in MECOH Lab has made my cultural preferences wider. Naomi: I always feel very grateful for having experience in MECOH Lab because finally I could learn many things about microbiology, DNA, and PCR. It’s so wonderful for meeting many people from different nations and could learn about trivial things about anything in this world, exchanging culture and experience will be always a thing I want to try by myself. I also want to thank to Kozo Sensei for giving me chance to join the lab and all the students that help me to learn. Hope MECOH Lab will be this joyful as the time goes by! Sidah: MECOH Lab will always be one of my great experiences. I learned many things in the laboratory, like being more disciplined, careful, and patient. I'm so happy to learn about DNA and molecular since it's a new topic for me. I want to thank Kozo Sensei, who allowed me to join his team, and Dan, who guided me patiently and explained the research in the Lab. People in MECOH Lab have warm hearts. I love the way they interact and support each other. Everyone is so gentle, supportive and passionate about doing the research.
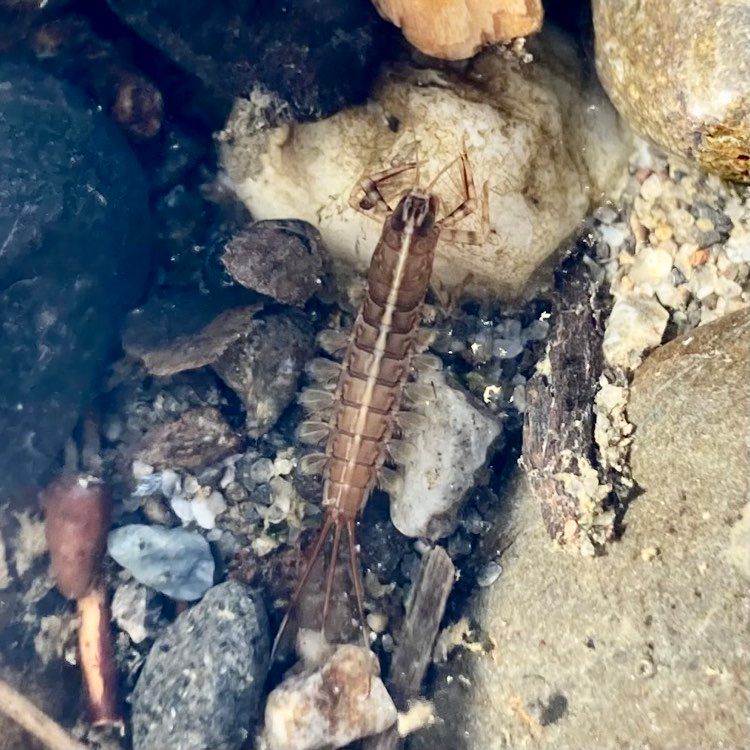
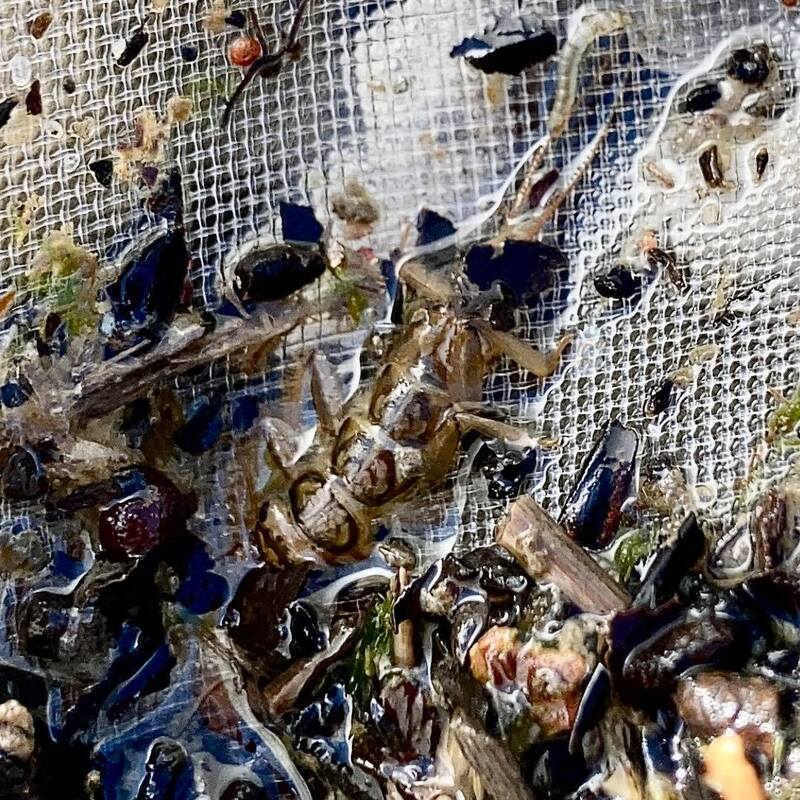
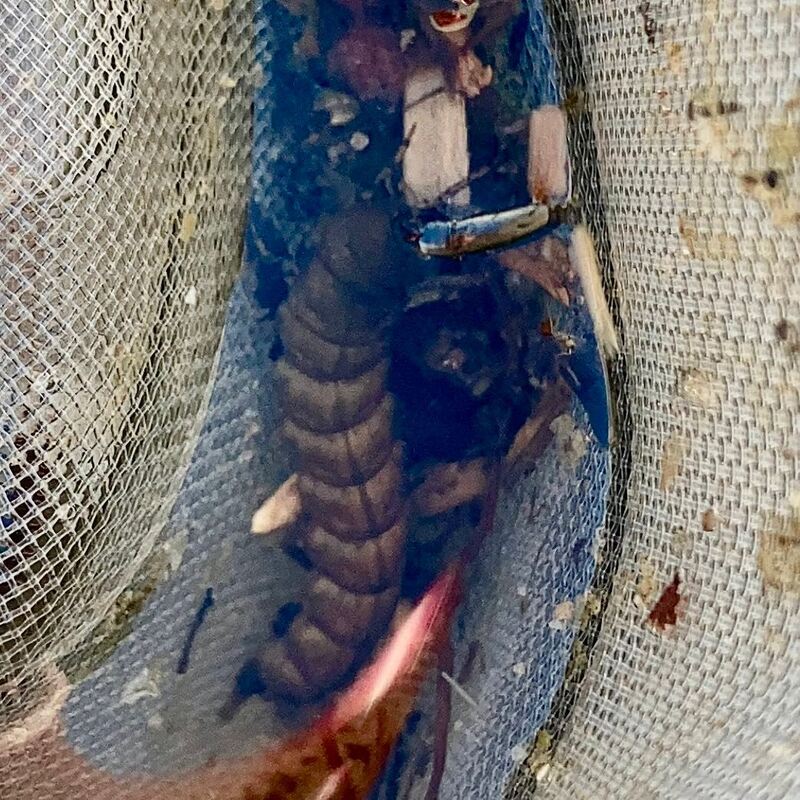
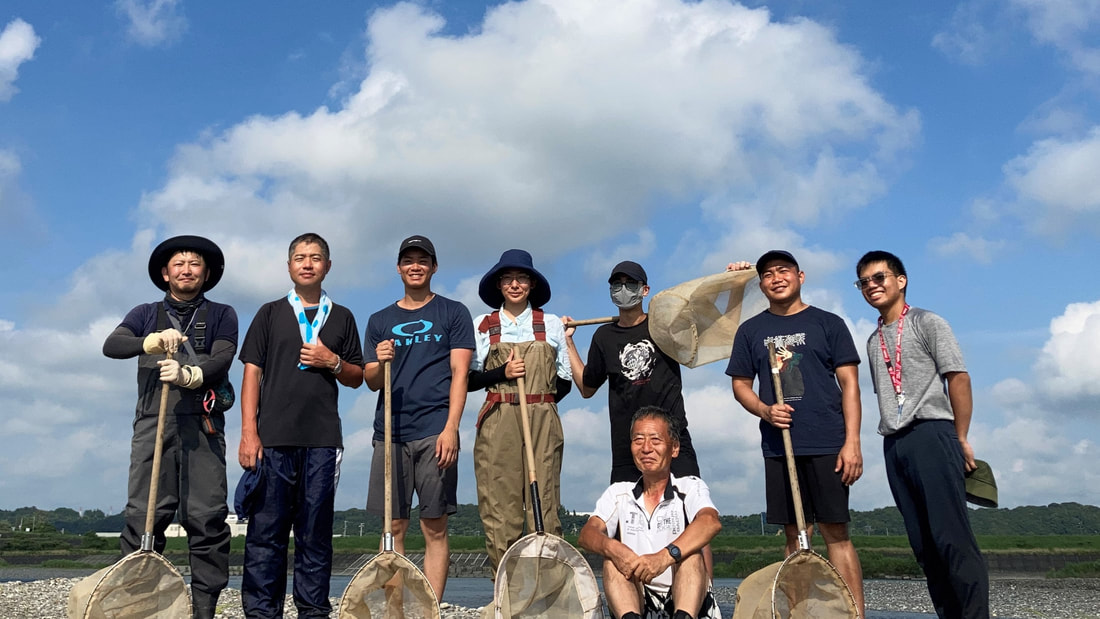
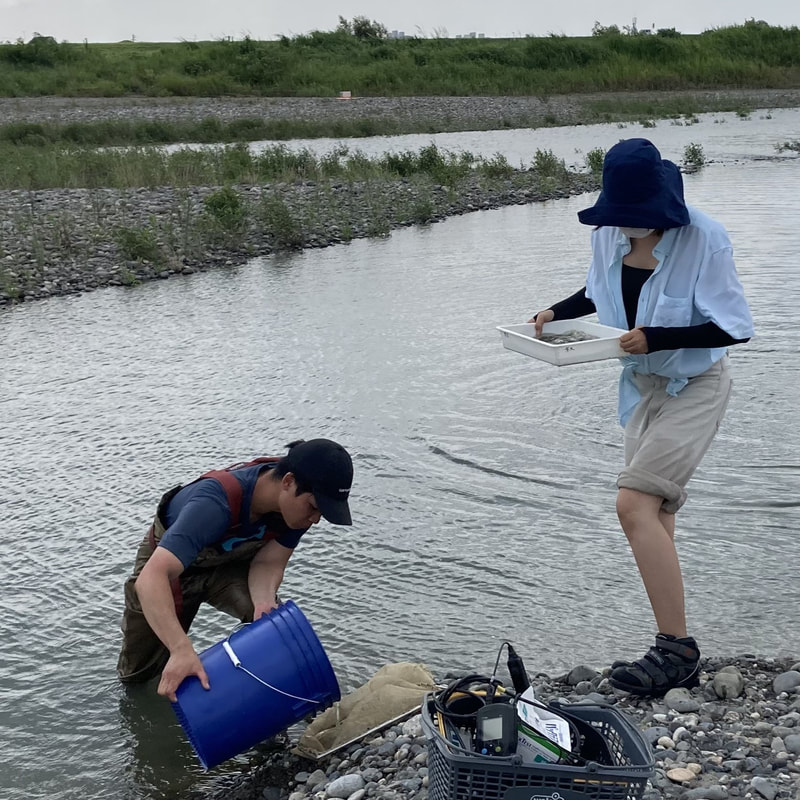
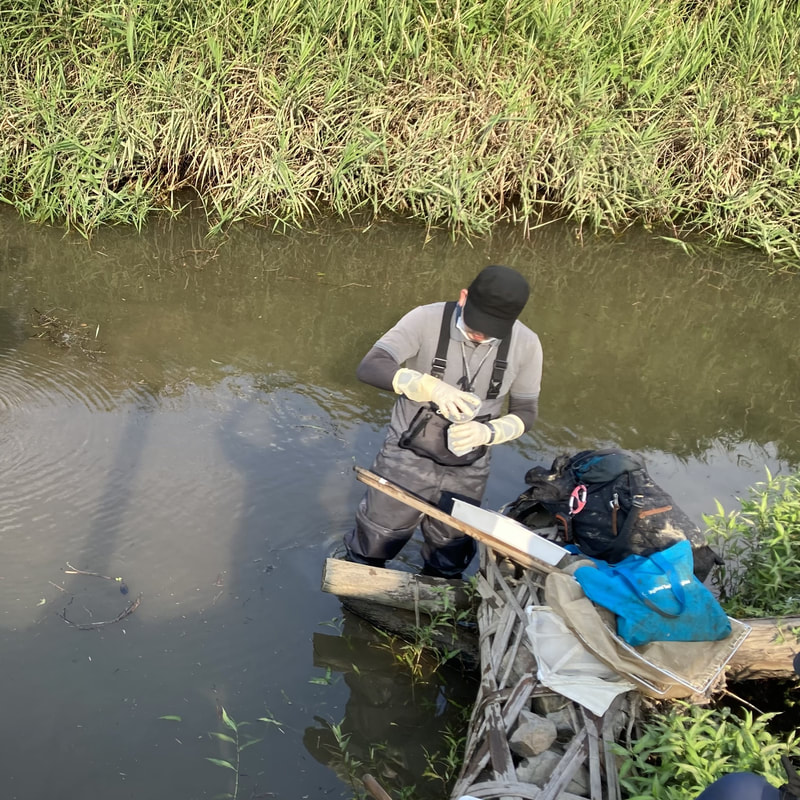
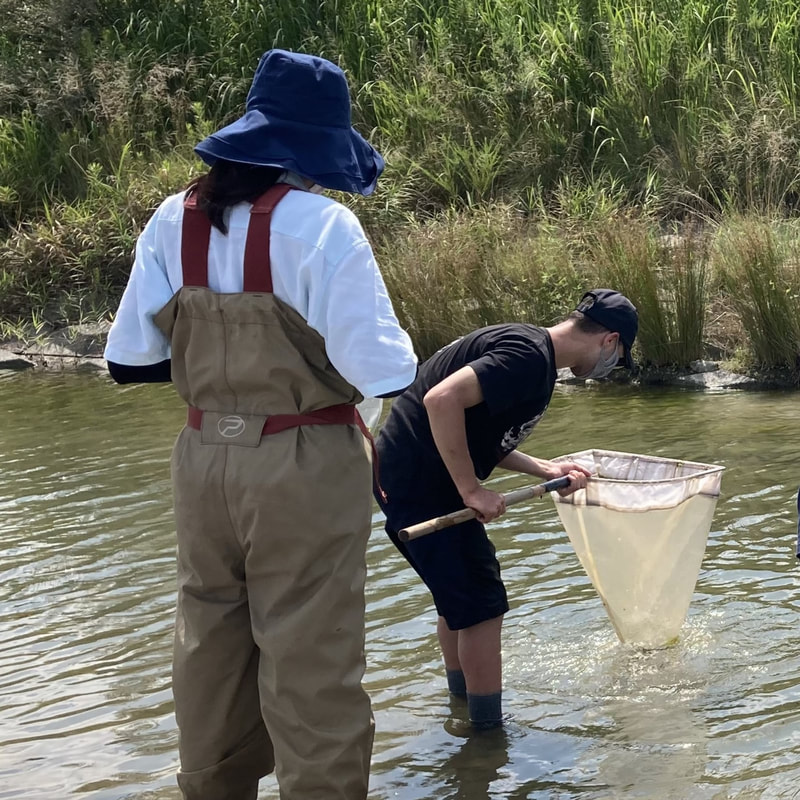
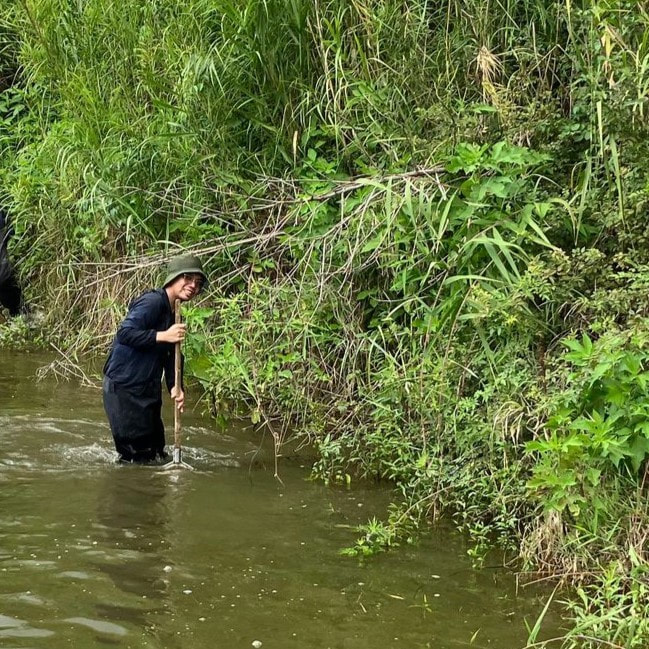
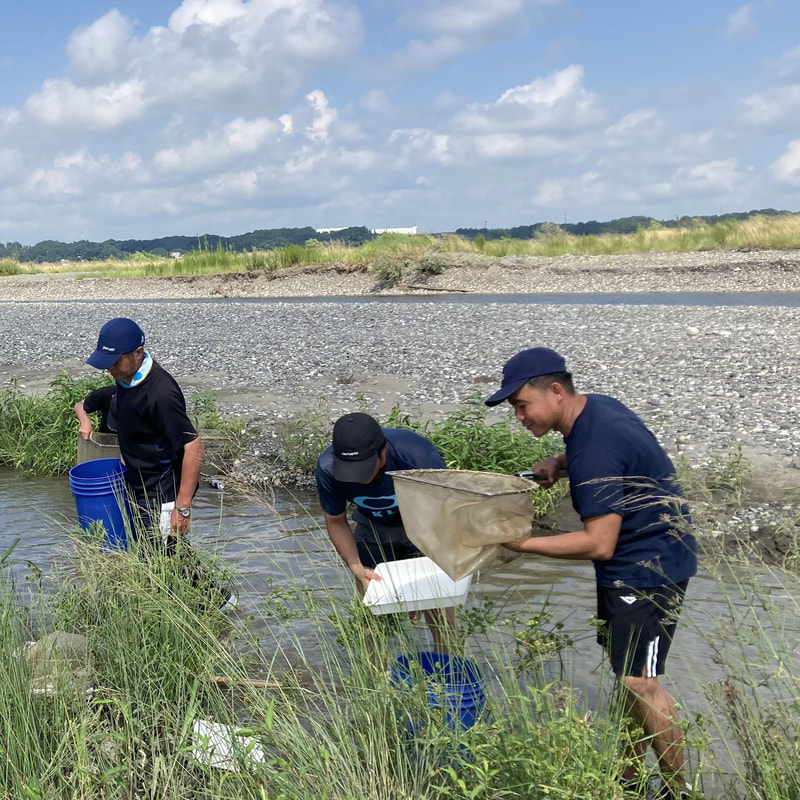
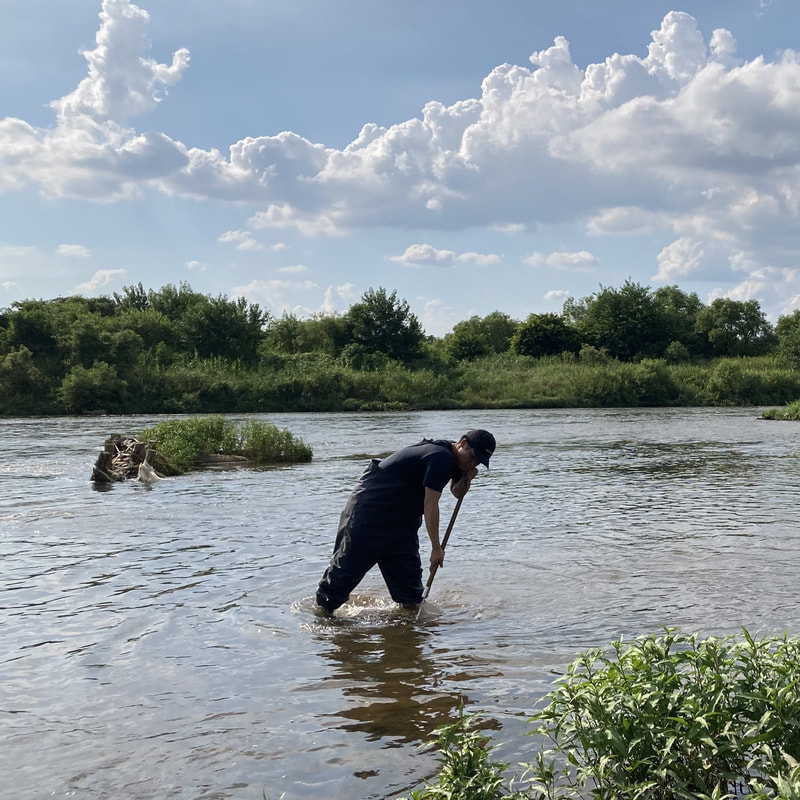
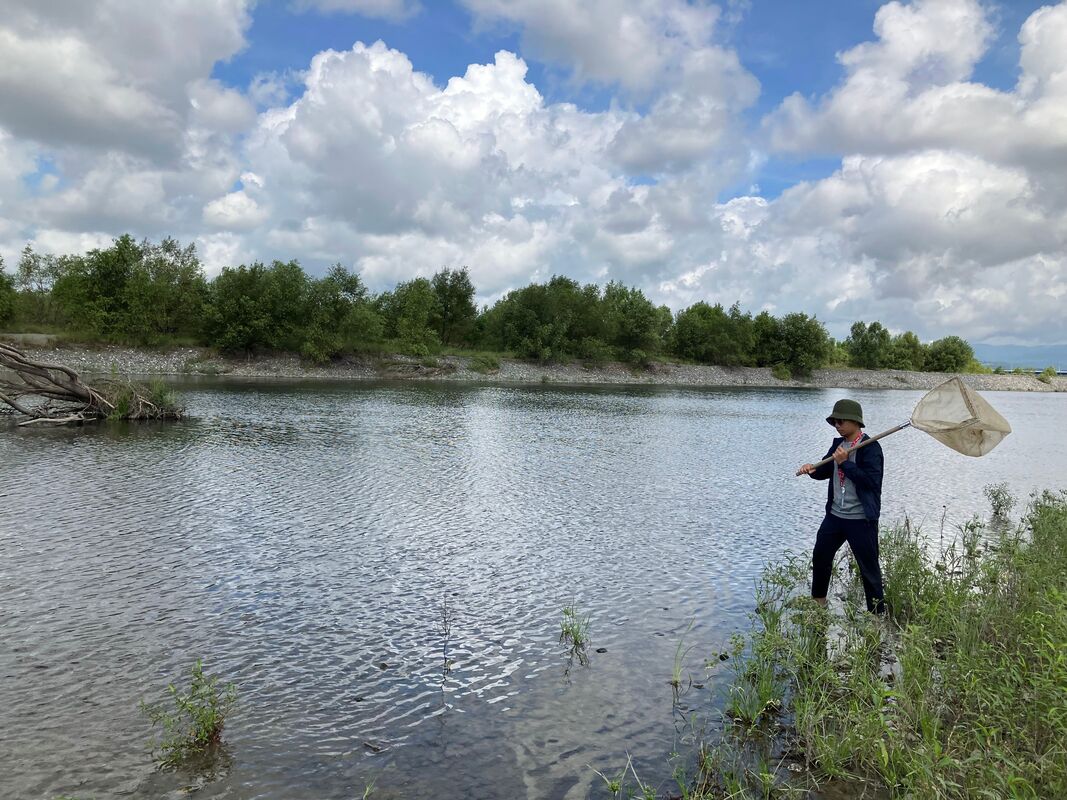
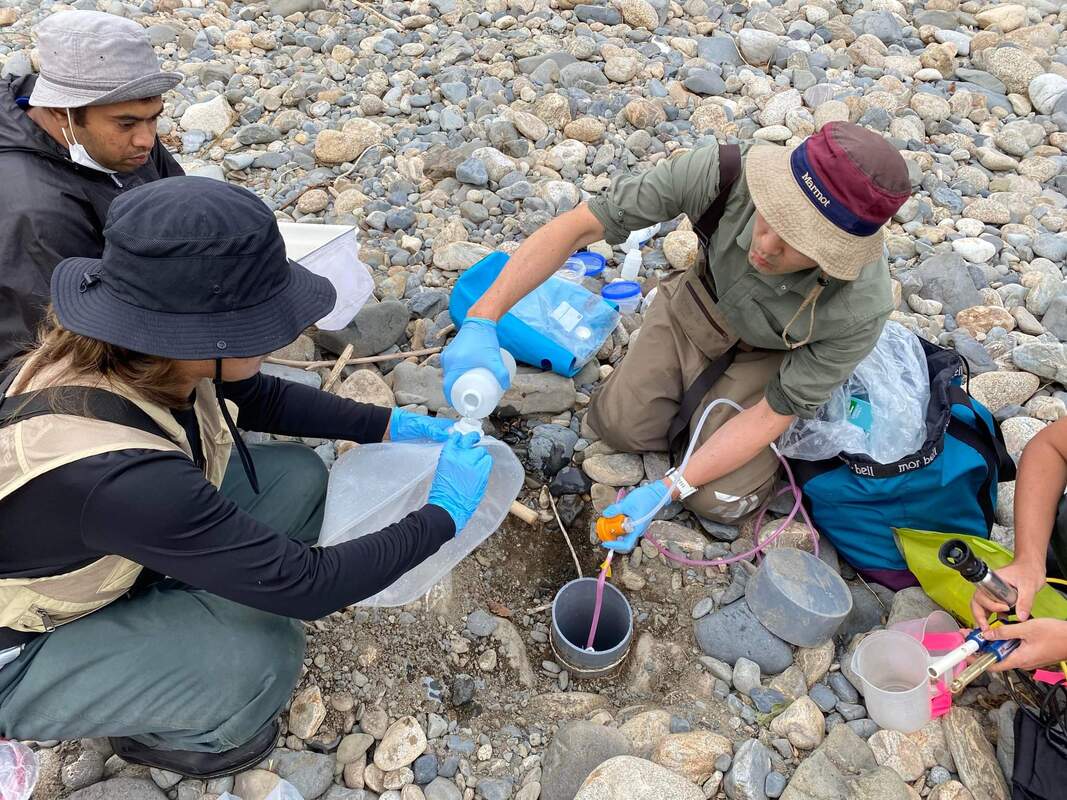
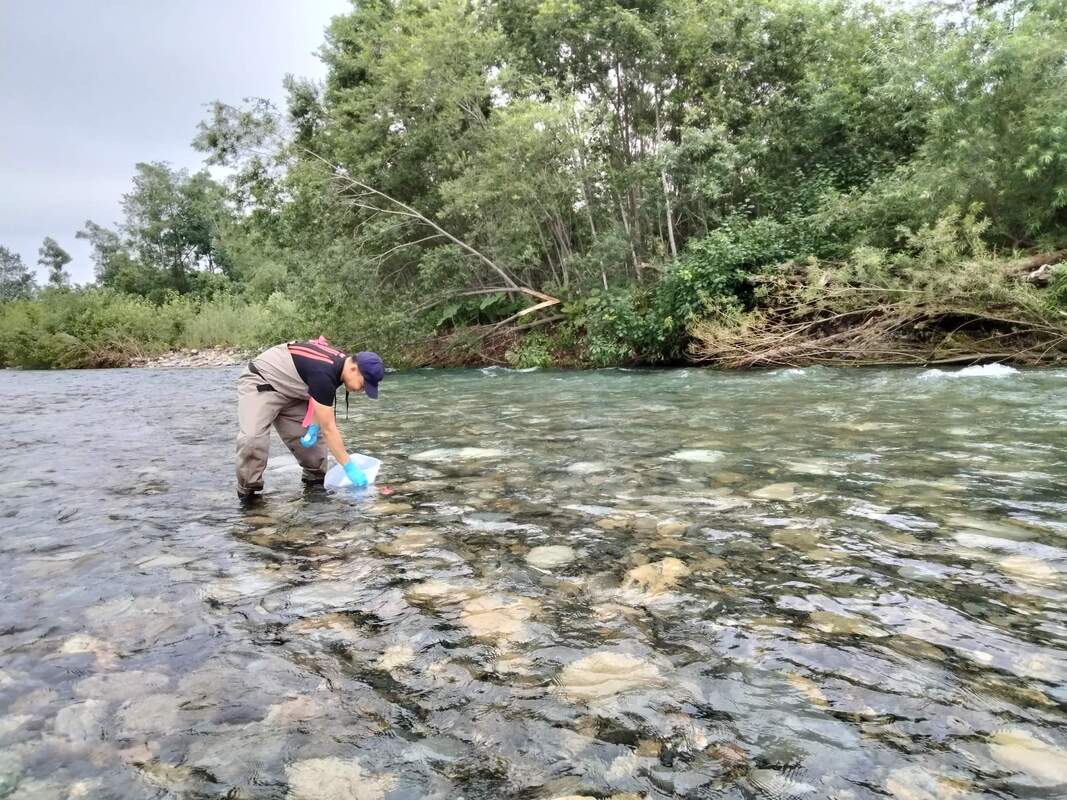
Vita: Firstly I came to Japan, I felt afraid because I felt constrained language when communicating in the Japanese language and also English. But, I can handle that because of support from Kozo Sensei and the others member in MECOH Lab, especially Ken and Ngure, who was very patient and explained well until I understood the research. This is my new experience studying DNA, and I am interested in this research. Thank you to the Japanese in MECOH laboratory, especially Shimada and Naoya, who accompany and provide insight into Matsuyama and Japan. Dr. Shinji Takahashi from Tohoku University, together with the Molecular Ecology and Health Laboratory members Mr. Arthien Pelingen (Researcher), Mr. Karim Clivaz (M2), and Mr. Tsubasa Moriya (B4), collected environmental DNA (eDNA) and macroinvertebrates from Tenryu River, Shizuoka Prefecture on November 9-11, 2022. This recently concluded field work also serves as a follow-up of the August collection trip this year (Read here: collaborative-fieldwork-of-three-universities-in-kizu-and-tenryu-rivers.html). Accordingly, we focused on (1) identifying Ayu (Plecoglossus altivelis altivelis) spawning sites and on (2) observing macroinvertebrate distribution in relation to the riverine spatial habitat diversity. On this three-day activity, we have successfully confirmed spawning of Ayu this season and positively identified different habitat types as we recorded various environmental parameters such as water temperature, pH, turbidity, etc. to be correlated in several macroinvertebrate studies.
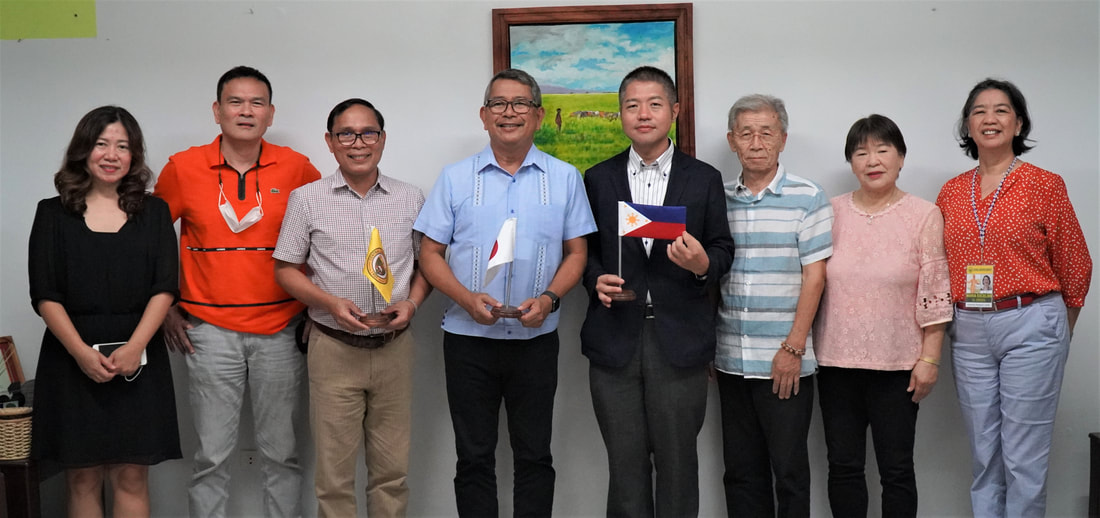
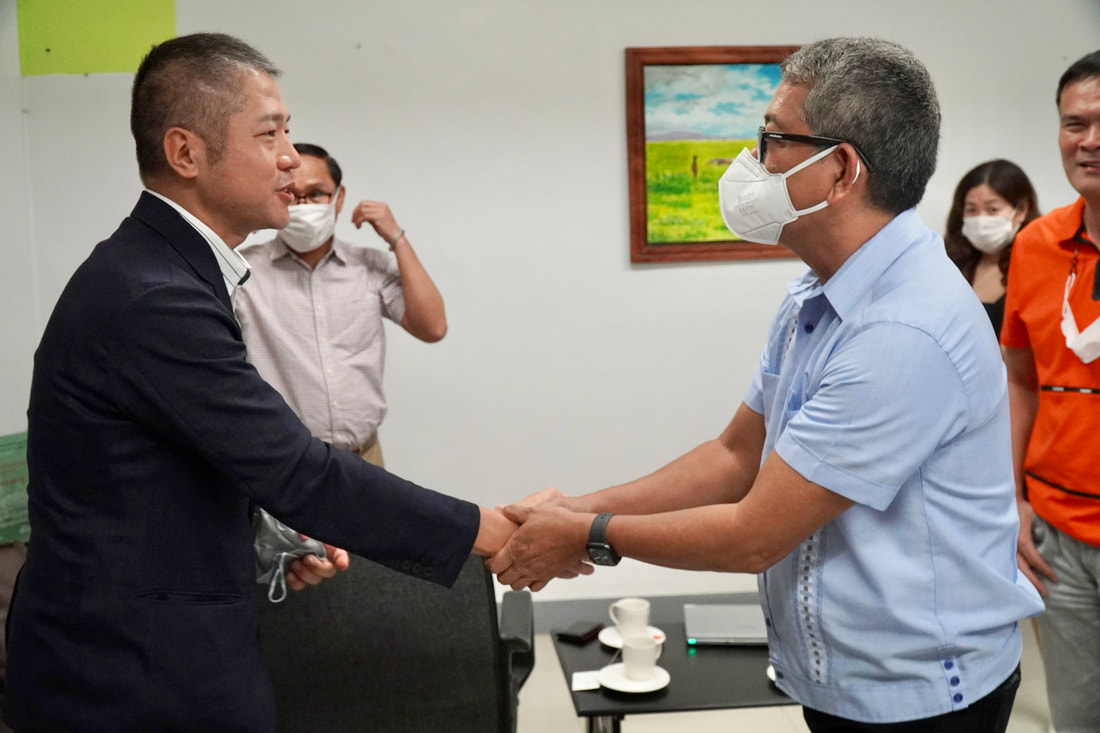
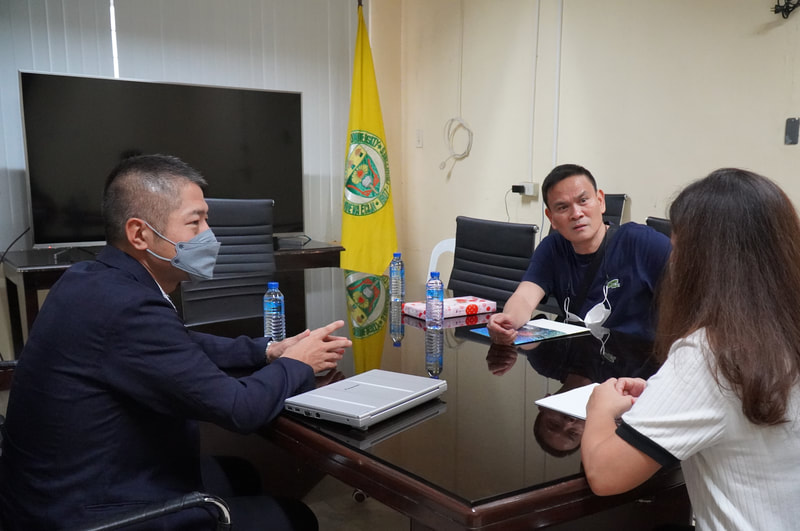
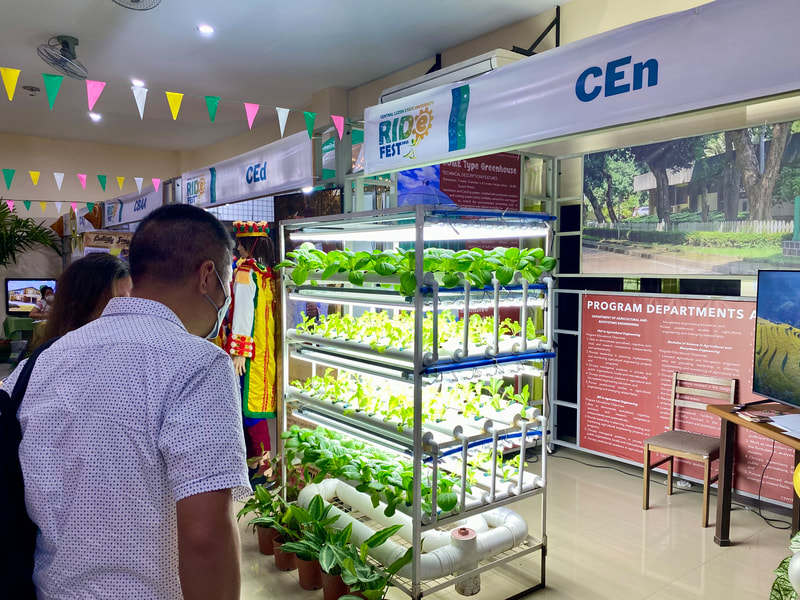
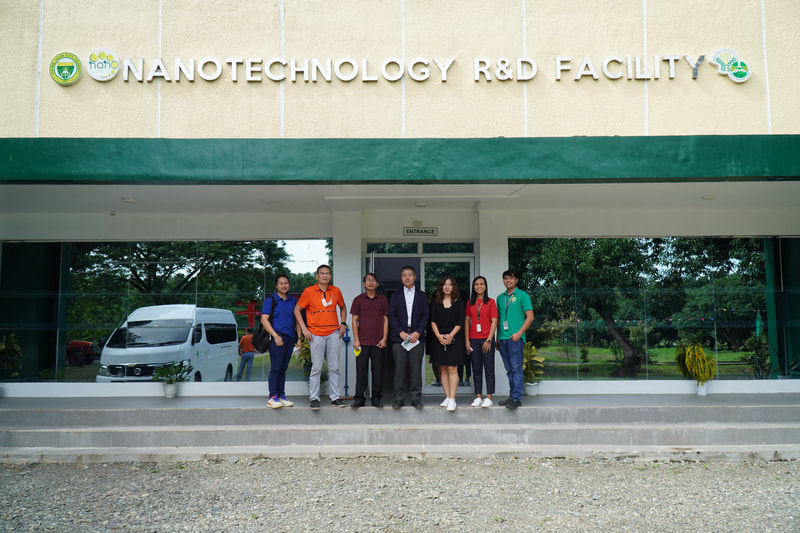
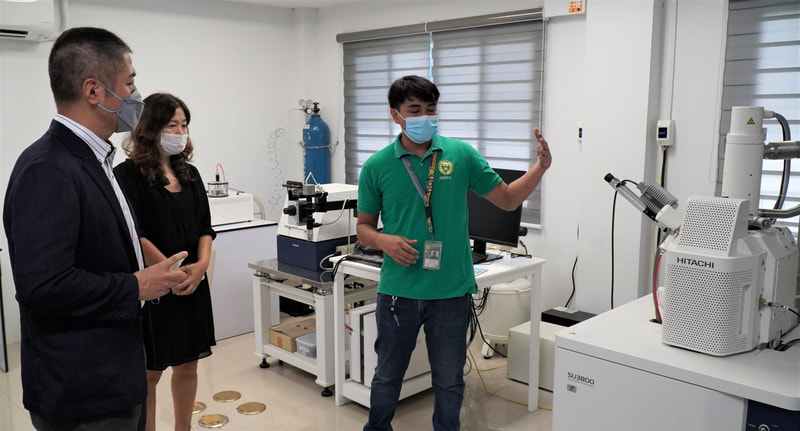
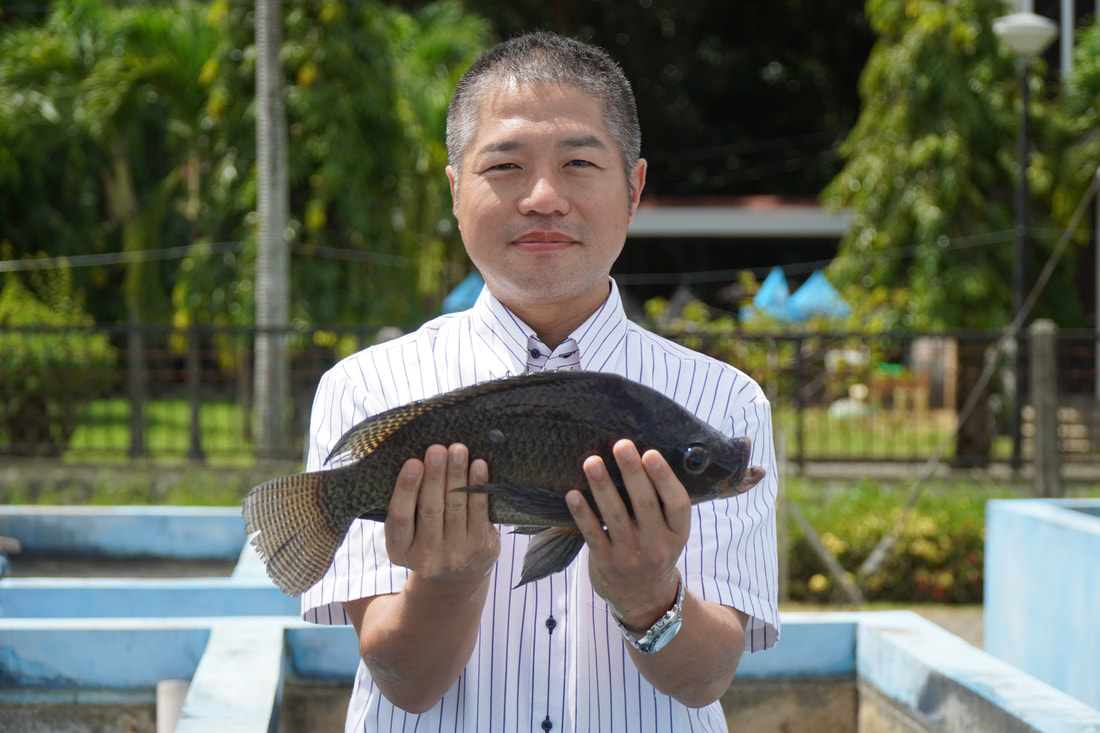
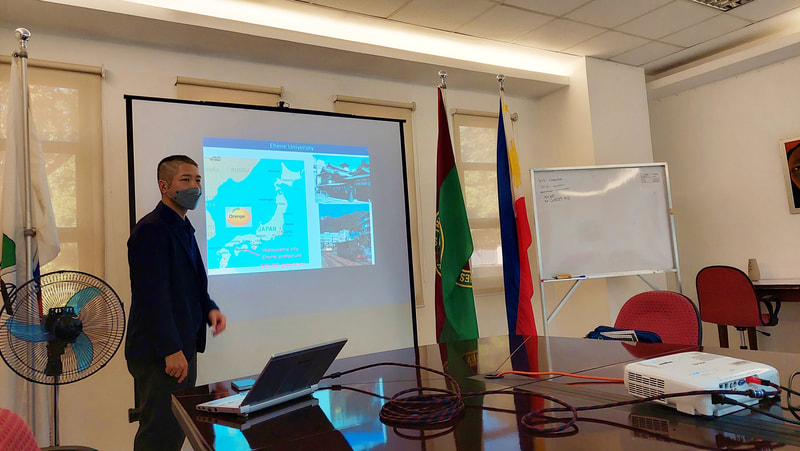
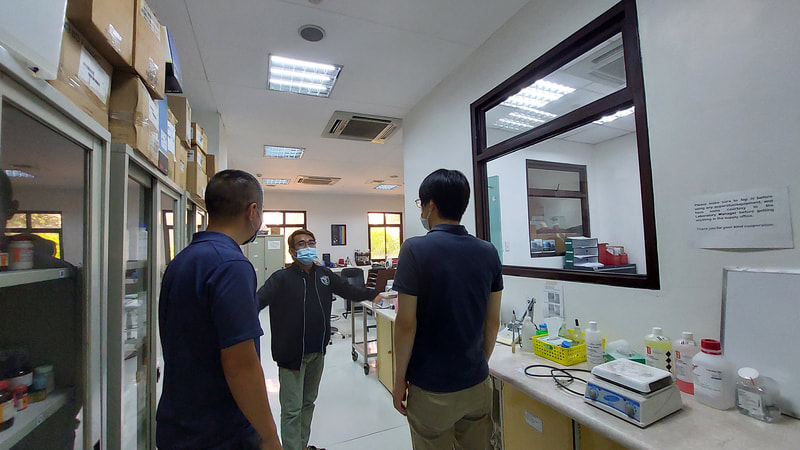
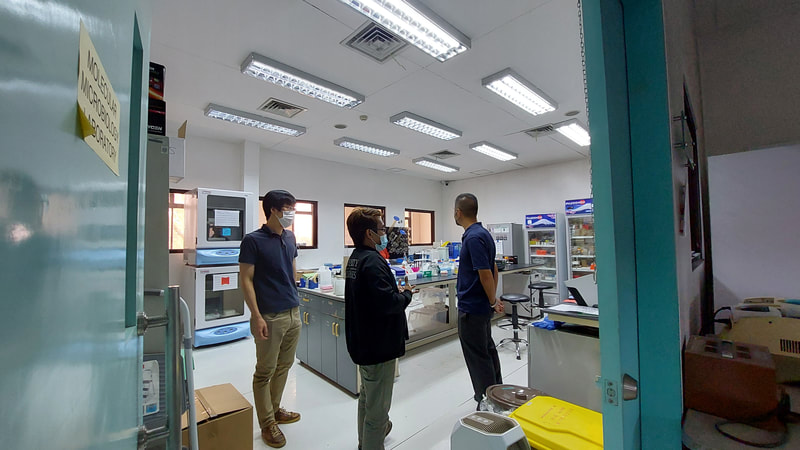
Ehime University-Center for Marine Environmental Studies (EU-CMES) represented by the head of the Molecular Ecology and Health (MEcoH) Laboratory, Dr. Kozo Watanabe and JSPS Postdoctoral Fellow Dr. Khristina Judan Cruz visited the Central Luzon State University, Science City of Munoz, Nueva Ecija, Philippines last October 12-13, 2022 for initial talks on the establishment of an EU satellite office in CLSU as well as the possibility of a cross-appointment of CLSU faculty members to EU. Dr. Watanabe and Dr. Judan Cruz discussed future EU-CLSU activities with CLSU President Dr. Edgar A. Orden, Vice President for Academic Affairs Dr. Renato G. Reyes, Dean of the College of Science Dr. Evaristo A. Abella, Dean of the College of Fisheries Dr. Ravelina R. Velasco and Director for Freshwater Aquaculture Center-CLSU Dr. Karl Marx A. Quiazon. A Memorandum of Understanding between the two universities was signed last January 22, 2022 to establish and strengthen International Collaboration for academic and research activities. Dr. Watanabe also visited several research projects and facilities in CLSU such as the CLSU Nanotechnology Facility and the Freshwater Aquaculture Center. The visit coincided with the CLSU Research Innovation and Development (RIDe) Festival that highlighted quality outputs of the university’s research and development projects. Current research collaboration between MEcoH and CLSU includes the genome-wide scan of genetically-improved tilapia particularly the FaST strain developed by CLSU, and integrating nanotechnology with bioactive compounds for anti-quorum sensing and anticancer evaluations.
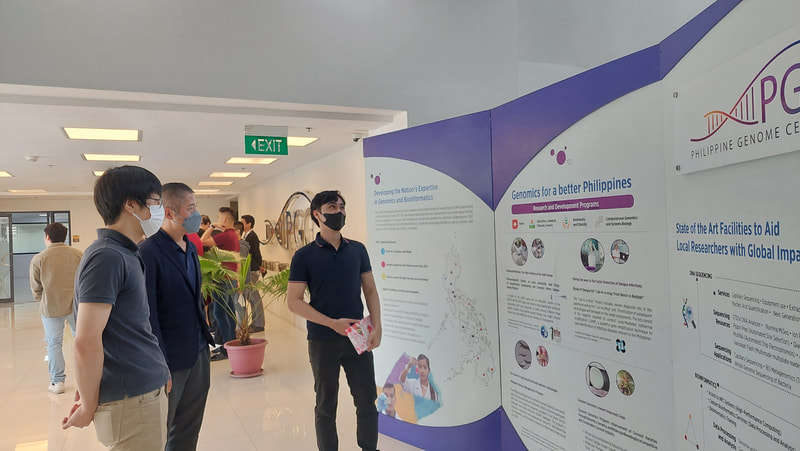
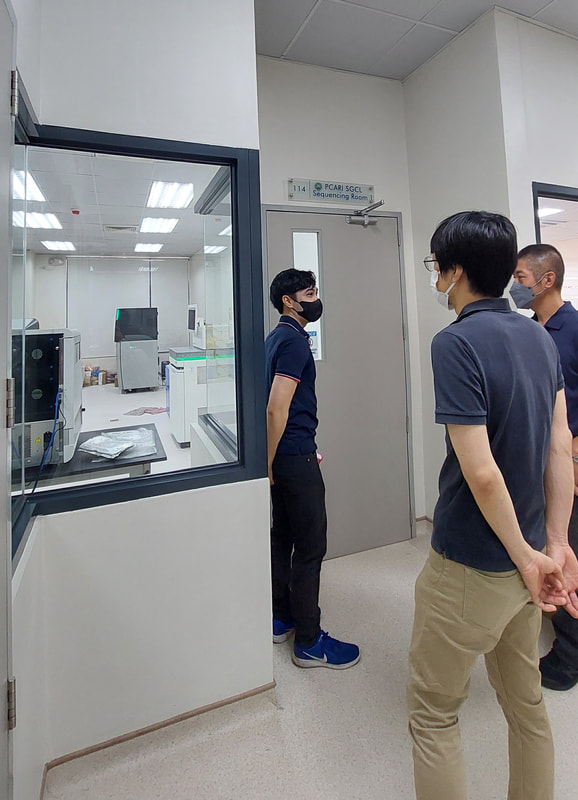
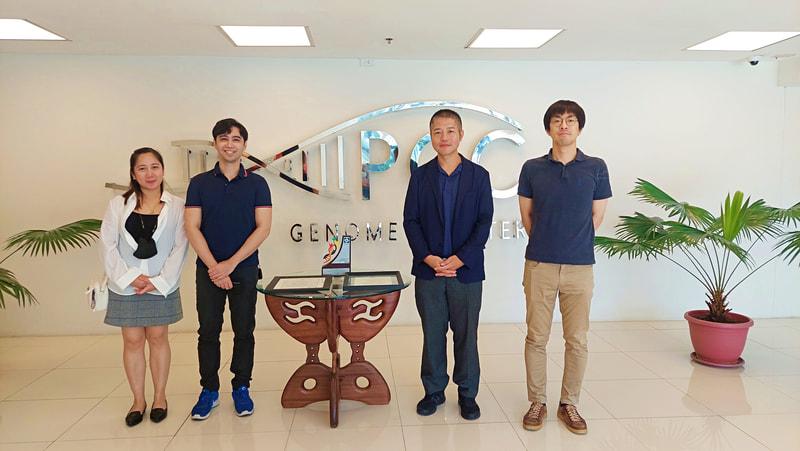
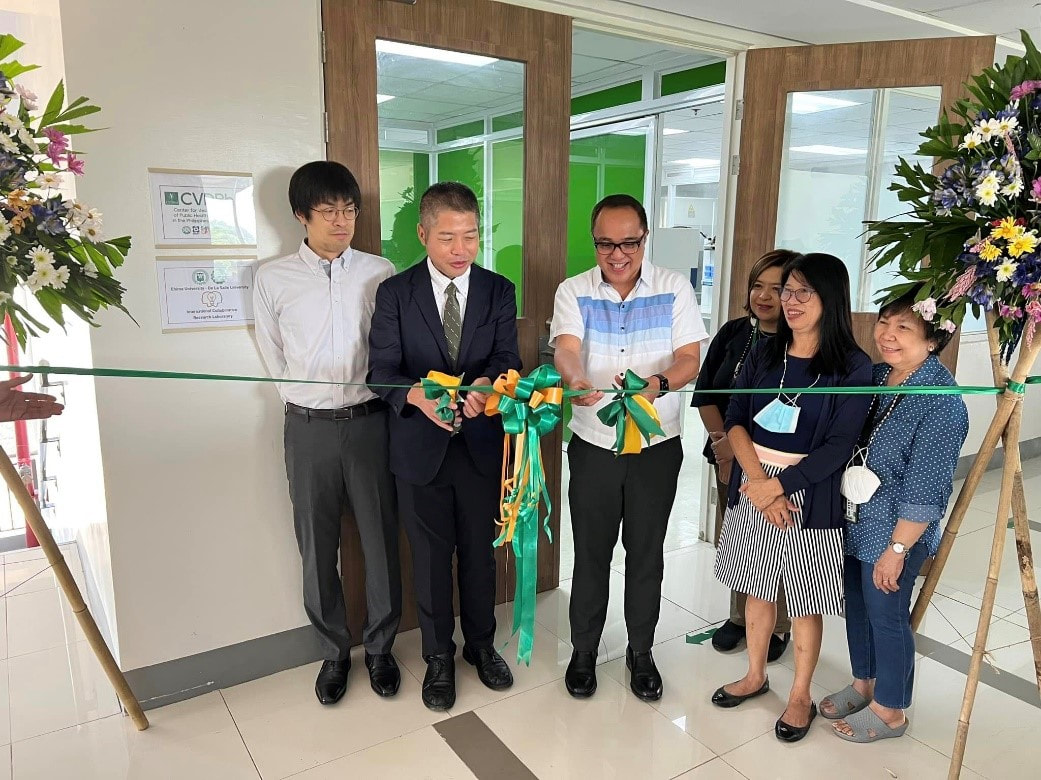
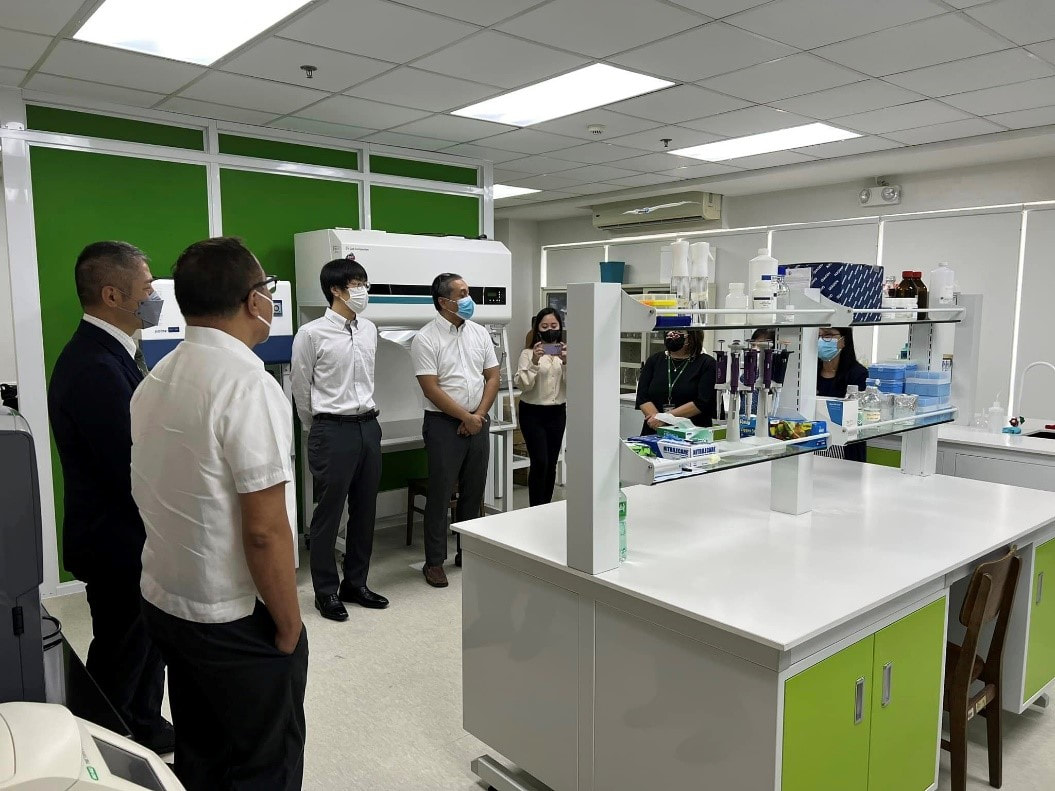
Potential Collaboration between Ehime University and NIMBB – University of the Philippines Diliman10/17/2022 Dr. Kozo Watanabe, head of the Molecular Ecology and Health (MEcoH) Laboratory, Ehime University (EU), together with Dr. Yasutsugu Suzuki (Project Associate Professor) and Ms. Irish Coleen Asin (PhD candidate) from the same laboratory visited the National Institute of Molecular Biology and Biotechnology (NIMBB), University of the Philippines Diliman (UP Diliman), Quezon City, Philippines last October 10, 2022. The representatives from Ehime University as well as Dr. Thaddeus Carvajal, Associate Professor at De La Salle University, met with Dr. Ma. Anita Bautista, Professor and Principal Investigator of the Functional Genomics Laboratory (FGL) of NIMBB-UP Diliman, for a talk on possible future collaboration between Ehime University and NIMBB. Also present in the meeting were Ms. Adria Lao (PhD candidate), Ms. Coleen Pangilinan (MSc candidate), and Mr. Aldwin Adiong (BSc candidate) of FGL-NIMBB who have presented their past and current research works on mosquito viral metagenomics and metatranscriptomics. Overall, this kick-off meeting provided an opportunity to build good rapport from both sides, and to set out and discuss various areas for collaborative work on mosquito research projects. Dr. Watanabe, Dr. Suzuki, and Ms. Asin have also been a given a tour of the facilities of NIMBB as well as the University of the Philippines System – Philippine Genome Center during this fruitful visit. Representatives from Ehime University (EU) and De La Salle University (DLSU) celebrated the official opening of the EU-DLSU International Collaborative Research Laboratory (ICRL) located at the George S.K. Ty Advanced Instrumentation Bldg, DLSU Laguna Campus on Tuesday morning, 11 October 2022. The ribbon-cutting ceremony was led by Dr. Kozo Watanabe, Professor of Molecular Ecology and Health (MEcoH) Laboratory at the Center for Marine Environmental Studies (CMES), EU, and Dr. Jonathan Dungca, Vice President for Laguna Campus and Dean of the College. Also present were Dr. Divina Amalin, Director of the Center for Natural Sciences and Environmental Research (CENSER), Drs. Ma. Luisa Enriquez, Mary Jane Flores, Thaddeus Carvajal, faculty members at the Department of Biology, DLSU, and members of the Biological Control Research Unit, CENSER. Also present are Dr. Yasutsugu Suzuki, Associate Professor of EU-CMES, and Ms. Irish Coleen Asin, PhD candidate from UE-CMES. The ceremony was followed by a tour of the facilities at the ICRL. The laboratory is equipped with advanced research infrastructure in line with its vision to “be the leading research unit dedicated to advancing, applying and facilitating high standards of research practice for Philippine and Japanese students, faculty and staff of both universities.” Under the mentorship of recognized scientists from EU and DLSU, the ICRL will provide a hub in conducting high-quality collaborative research between Filipino and Japanese students, faculty, and staff. A seminar session was also conducted wherein Dr. Kozo Watanabe first discussed the functions and roles of the EU-DLSU ICRL. Dr. Thaddeus Carvajal and Dr. Yasutsugu Suzuki also delivered their talks on “The Importance of Studying Vector Biology & Ecology of Mosquito-borne Diseases for Designing Appropriate Vector Control Strategies” and “Understanding Mosquito-virus Interactions Toward Vector Control”, respectively. The seminar was livestreamed and was attended by participants online. The ICRL is a product of a long standing cooperation between the two universities that started in 2016. In October 21, 2021, the Memorandum of Agreement was signed by Dr. Nishina Hiroshige, President of EU, and Br. Bernard Oca, President of DLSU, to start the establishment of ICRL (then known as the Ehime University Satellite Office Philippines). Currently, the ICRL implements several privately and publicly funded research projects, and houses the Center for Vector of Diseases of Public Health Importance in the Philippines for Region IV, a niche center under the DOST Science for Change Program.
On 17th - 22nd September 2022, Professor Kozo Watanabe visited two universities in Bangladesh -Dhaka University, at Dhaka city and Mawlana Bhashani Science and Technology University (MBSTU), at Tangail city. Professor Watanabe attended in several meetings and seminars organized by both universities and discussed about the possible way of future collaborations. Mohamad Mosleh Uddin, a PhD student of MECoH lab went Bangladesh (22nd August -28th September, 2022) for his sampling purpose. He collected mosquito samples from the Dhaka City area. The first face-to-face workshop for the JSPS core to core program was held successfully last 2022 September 15 at Ehime University. Several participants from the Center for Marine Environmental Studies (CMES) and Proteo-Science center joined the workshop. Interesting topics were discussed by the invited speakers. Dr. Watanabe opened the symposium followed by the special lecture by Dr. Hann from National University of Singapore. Dr. Hann gave a glimpse of the projects from his laboratory one of which is on the recent advances in antiviral strategies against medically important positive-sense RNA viruses. Dr. Kuwata from Okayama University of Science gave an interesting talk about the status of mosquito-borne arboviruses in Ehime. Dr. Hirano from Nagasaki University shared his research on Crimean-Congo hemorrhagic fever virus, which is BSL4 pathogen. Dr. Suzuki from Osaka Medical and Pharmaceutical University presented novel virus titration assays and screening for antivirals. From Ehime University, Dr. Takahashi and Dr. Suzuki shared their current studies on host-virus interactions. In summary, the symposium successfully presented diverse topics in virology paving the way for future collaborations.
A workshop on emerging and re-emerging viruses will be held on 15 September 2022 from 14:30. Six speakers will cover diverse topics on virology not limited to antiviral strategies against medically important positive-sense RNA viruses and host-virus interactions.
On August 7-10, 2022, the members of MEcoH Lab led by Prof. Kozo, together with Dan and Art (Research students), and Tsubasa (B4) joined the Kyoto University team headed by Prof. Takemon, along with Liu (D1), Nishimura (M1) and Mojar (M1) as well as Dr. Takahashi from Tohoku University in a 4-day collection trip in Kizu and Tenryu Rivers. Along these large rivers, different habitat types were identified, inspected, and compared with one another. Despite the scorching heat, the fieldwork was successful as samples for several projects were collected including macroinvertebrates and eDNA.
At the start of August 2022, Art from the Philippines joined the MEcoH Lab as the newest PhD student. Emerging from a rigorous taxonomic training of Philippine macroinvertebrates during his Master’s in the Ateneo de Manila University (AdMU), Art will explore combined ecological and genomic approaches in studying the stream macroinvertebrates i.e. how the abiotic factors and genetic similarities and/or differences play into their roles in the river dynamics. With these in sight, he envisions to contribute into an interdisciplinary understanding of addressing the changes in current environmental patterns using macroinvertebrates as models.
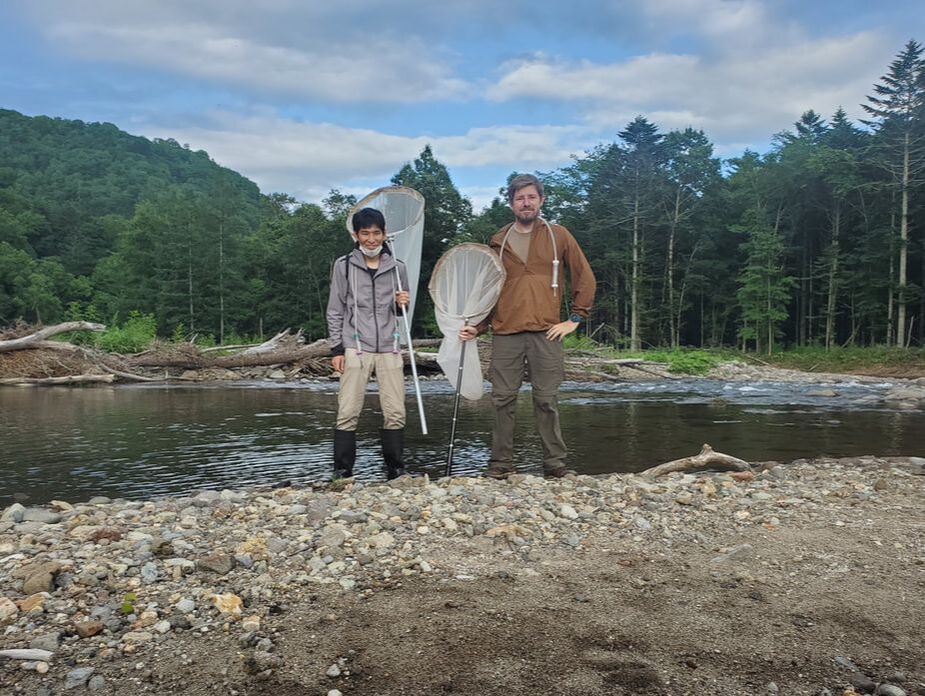
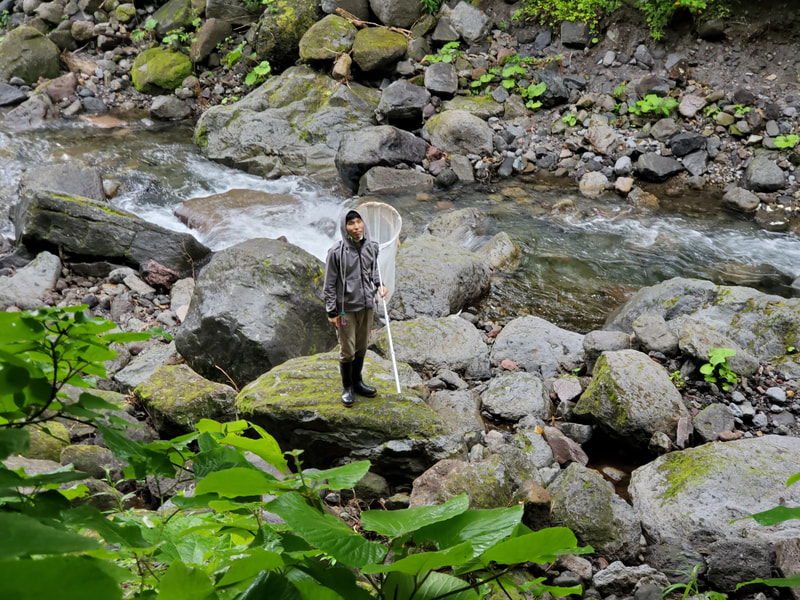
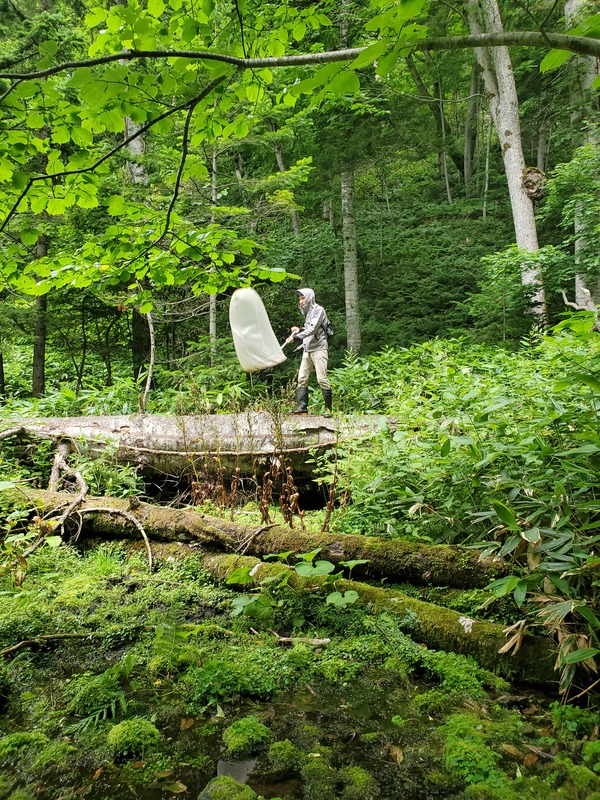
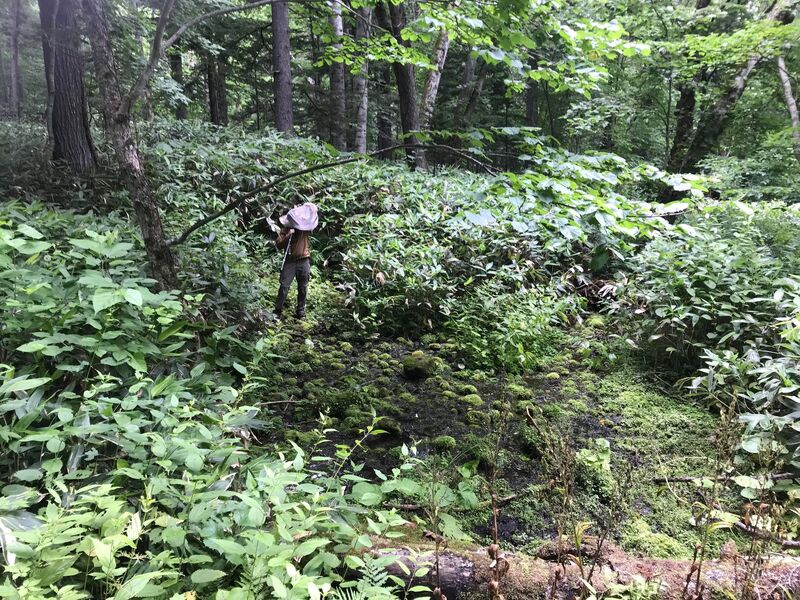
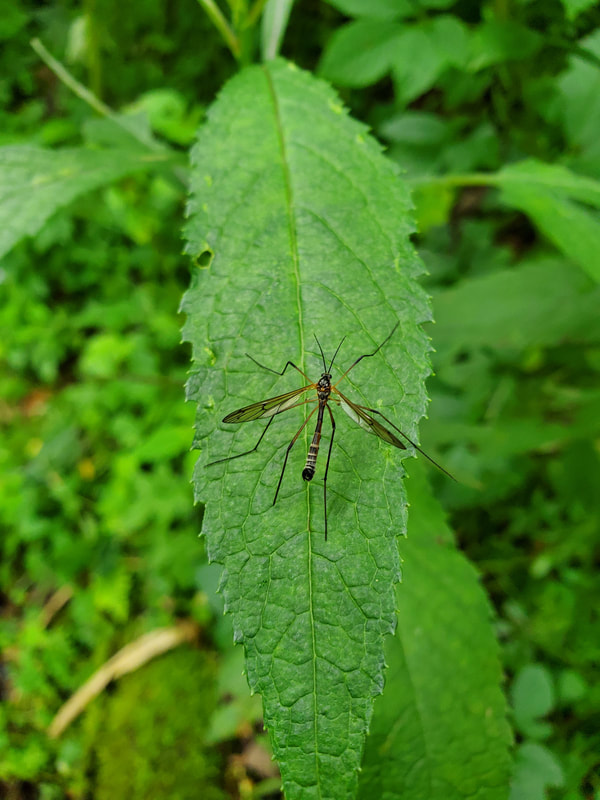
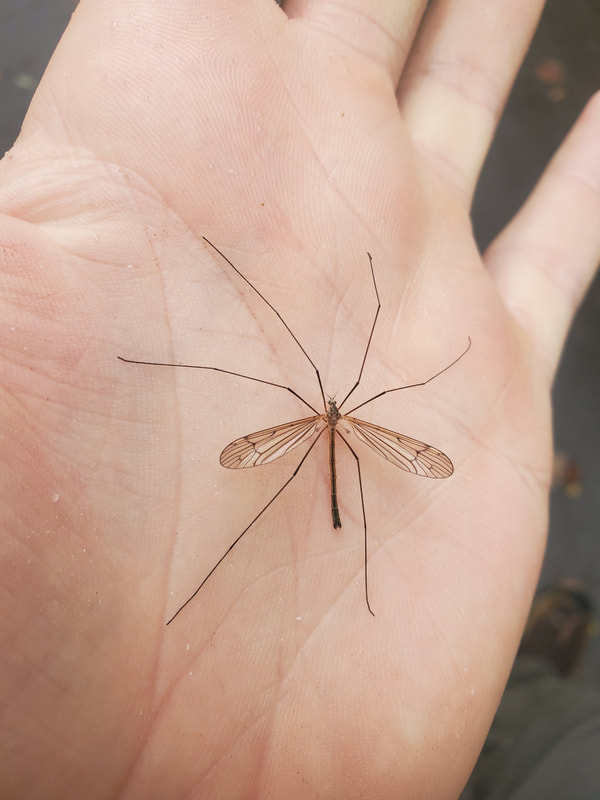
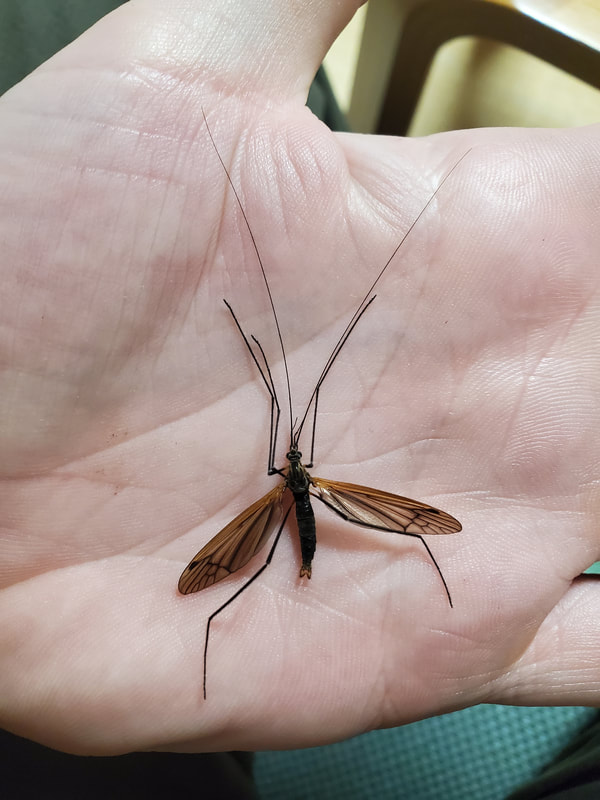
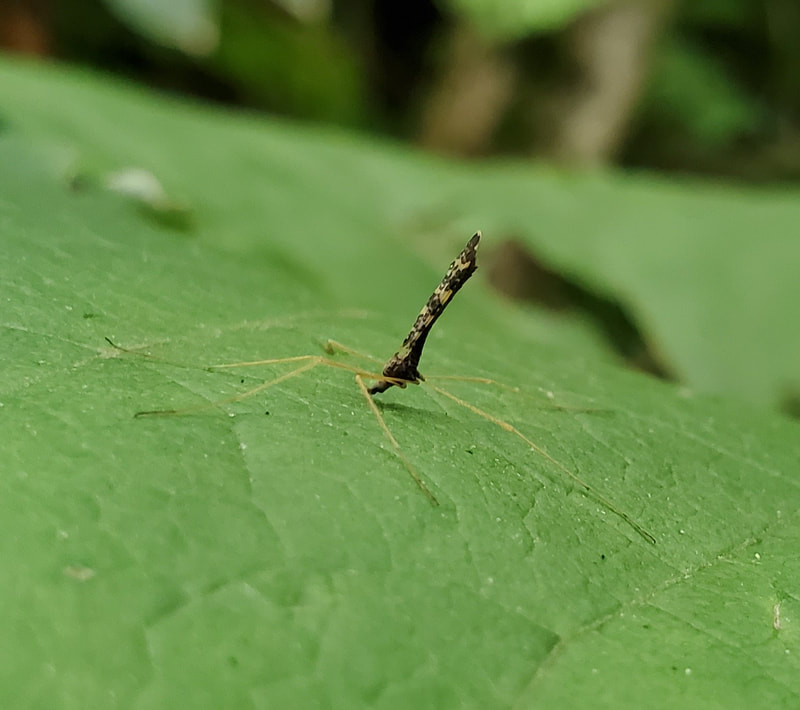
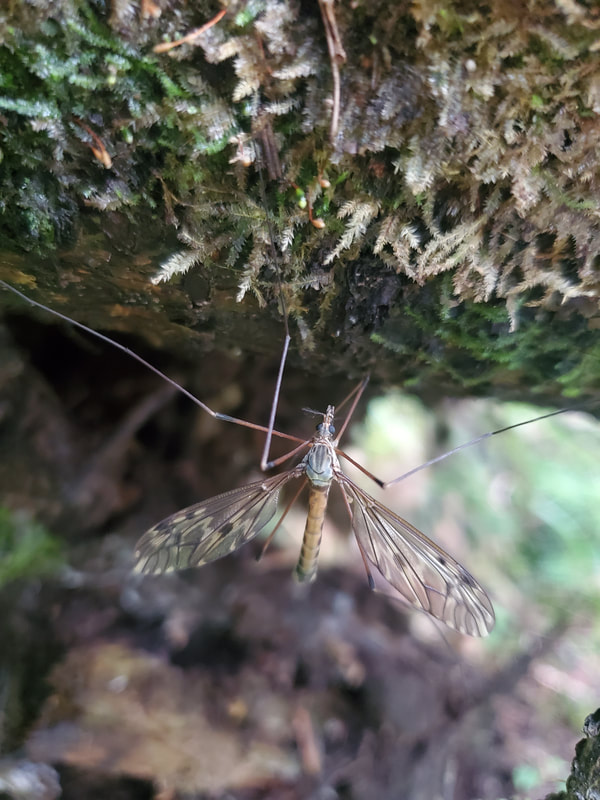
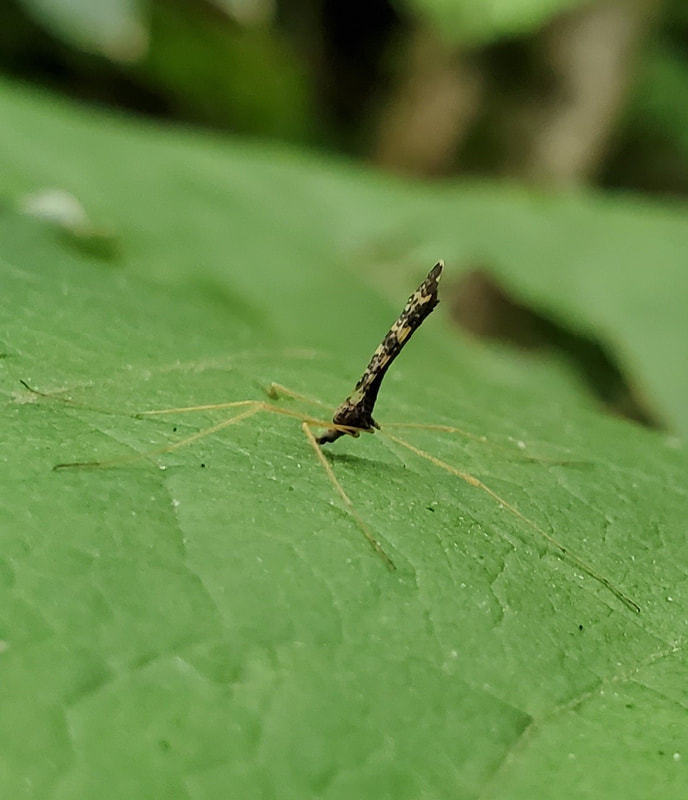
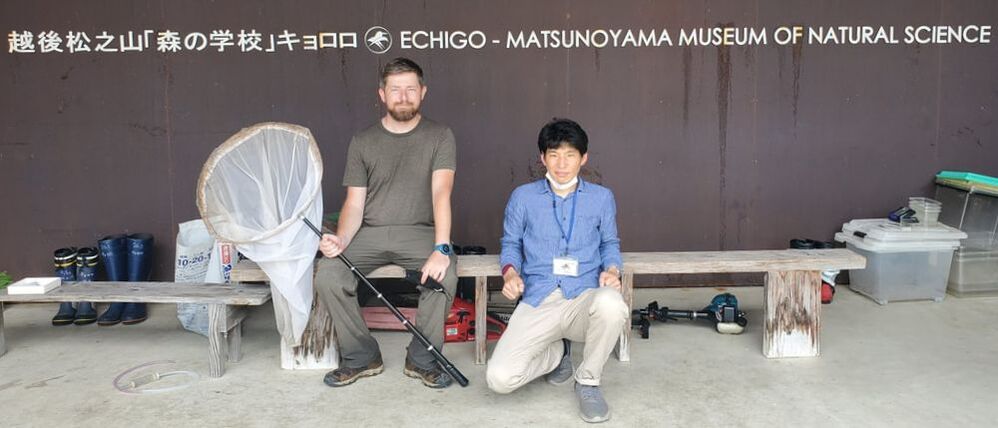
Dr. Levente-Péter Kolcsár (JSPS postdoctoral fellow) went on a field collection trip to Hokkaido between 18-29 July. Dr. Daichi Kato from Echigo-Matsunoyama Museum of Natural Science, Matsunoyama, Niigata Pref. also joined the field collection between 19-22 July. Aims and preliminary results of the trip: 1. Collecting additional specimens from two species new to science, which first time collected during another field trip in Hokkaido, in 2019. Both new species are only known to be found in 2 locations in 2019. During a recent field trip the new species were also found in additional 5 localities. 2. Collecting rare or poorly known crane fly species for future taxonomic studies. 3. A general survey of Hokkaido’s crane fly fauna. Due to the special geographic position of Hokkaido Island we suspect the presence of several species which are known from continental Asia, North-America and Kuril Island, but not yet reported from Hokkaido. In total dr. Kolcsár and dr. Kato collected around 1500 specimens from 30 collection sites. Some poorly known species collected during the field trip.
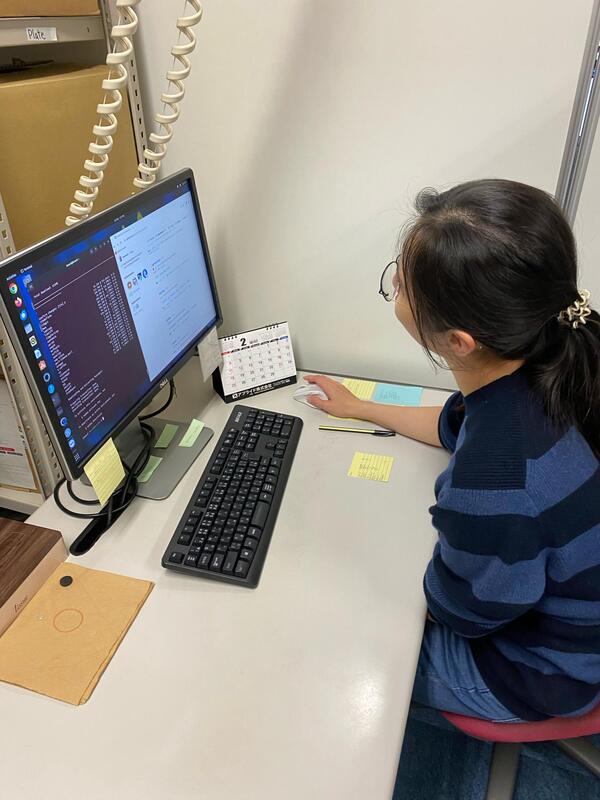
I am Maria Angenica F. Regilme, or Nica for short, I am currently a PhD student in the MECOH laboratory. I joined the laboratory as a Research student in September 2016 as a Japanese government scholar and started my PhD program from 2018 April until present. My PhD dissertation’s main goal is to use a wholistic approach of studying the relationship between the vector, host, microbiome including the pathogen and the environment to create a substantial information for an effective vector borne disease (VBDs) control. For my dissertation, I focused on the top VBDs: mosquito-borne diseases focusing on dengue mosquitoes, Aedes aegypti and Aedes albopictus and tick-borne diseases vectors, Ixodes ovatus and Haemaphysalis flava. The overall objective of my dissertation is to know the gene flow pattern among arthropod vector (e.g. mosquitoes and ticks) populations can be influenced by environmental factors (e.g.roads and altitude) and the hosts affecting the pathogen transmission. The presentation is for 45 minutes followed by question and answer portion from the audience for 15 minutes and a close meeting with the thesis panelist: Kozo sensei, Kitamura sensei and Miyake sensei. Finally, after more than 3 years I was able to present my thesis dissertation last 18 August, it was a hybrid style of thesis presentation wherein the thesis panelist are online via zoom while I present my thesis with my labmates in the seminar room. Before my presentation, I was nervous but I told myself to do my best because this is my only chance to get a PhD degree and to share the importance of my research to controlling vector-borne diseases. After the presentation and the question and answer portion, tears of joy fell into my eyes, I could only thank God, my family and friends, MECOH lab members (former and present), co-authors, collaborators, funding agencies and everyone who helped me in this PhD journey and most especially Kozo sensei for the support and guidance, without it, I will not be able to present my dissertation. I still don’t know the results of my thesis dissertation if I can be able to get the PhD degree. I can only hope that through God’s will I can get the degree and still continue doing research after PhD that can have an impact on improving the lives of people for example, research about tropical diseases. My heart is full of gratitude to everyone who have helped me, almost 6 years of PhD journey is one of the most memorable moments in my life that shaped me to become a future scientist and taught me many learnings in my life. Again, thank you!
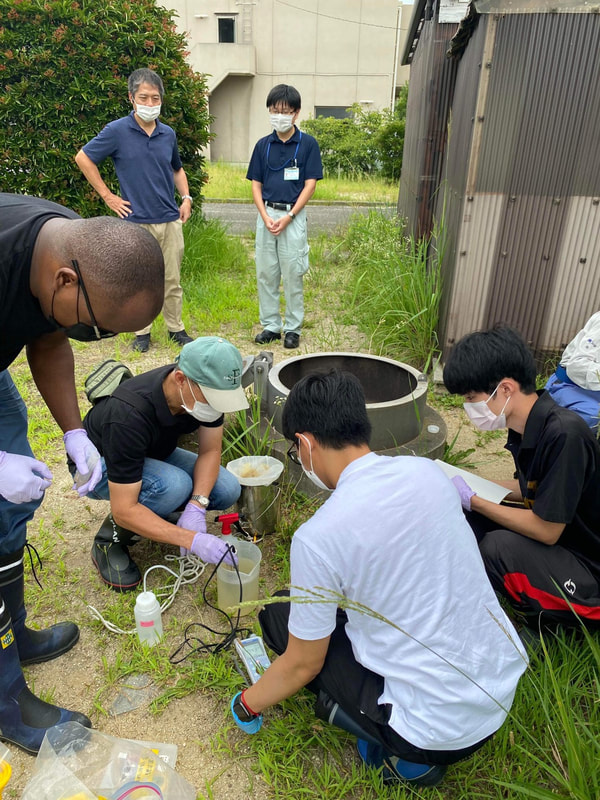
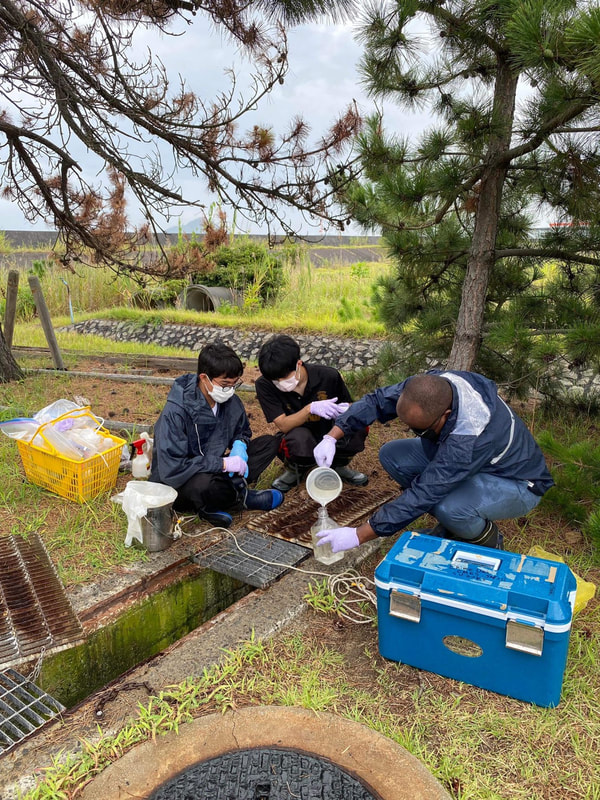
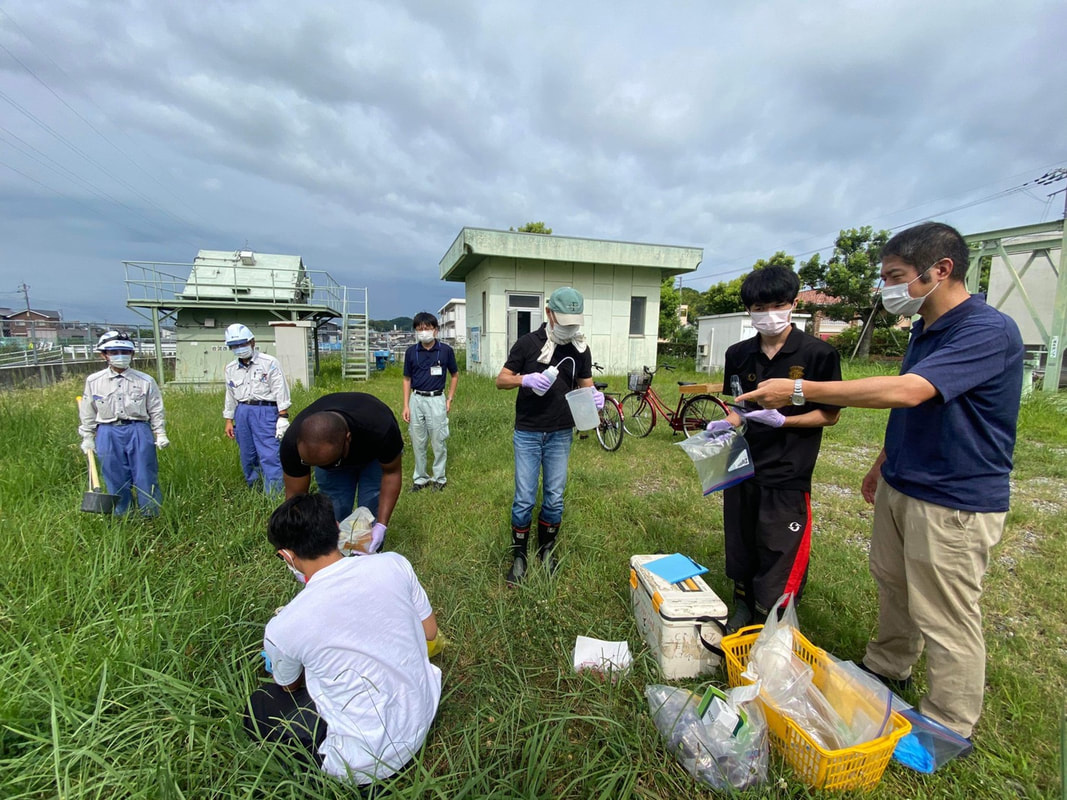
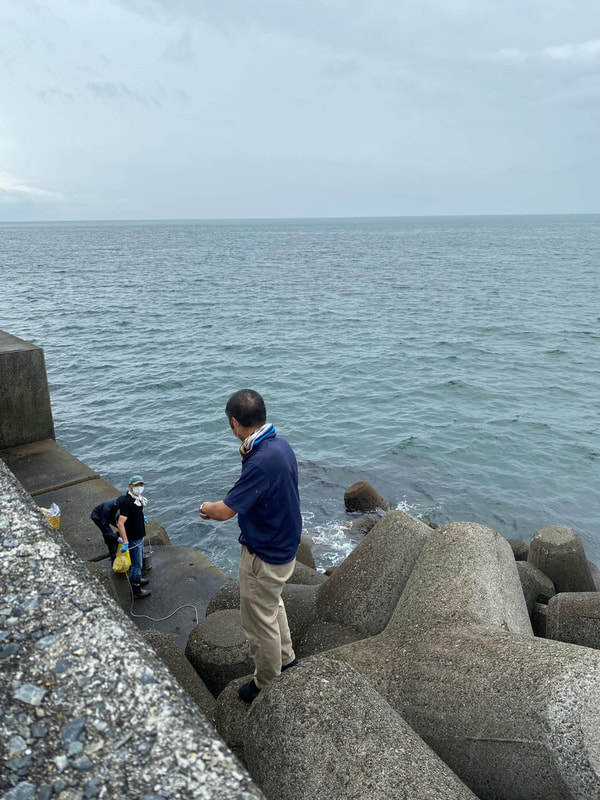
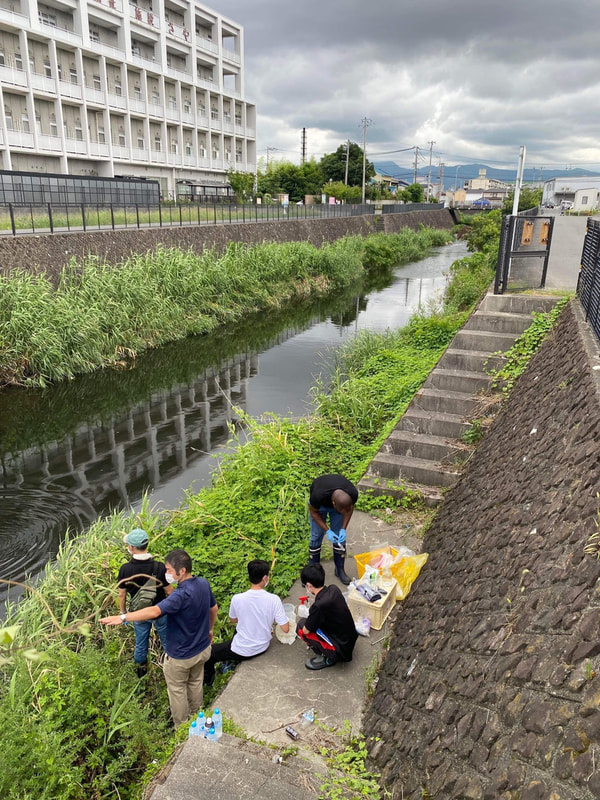
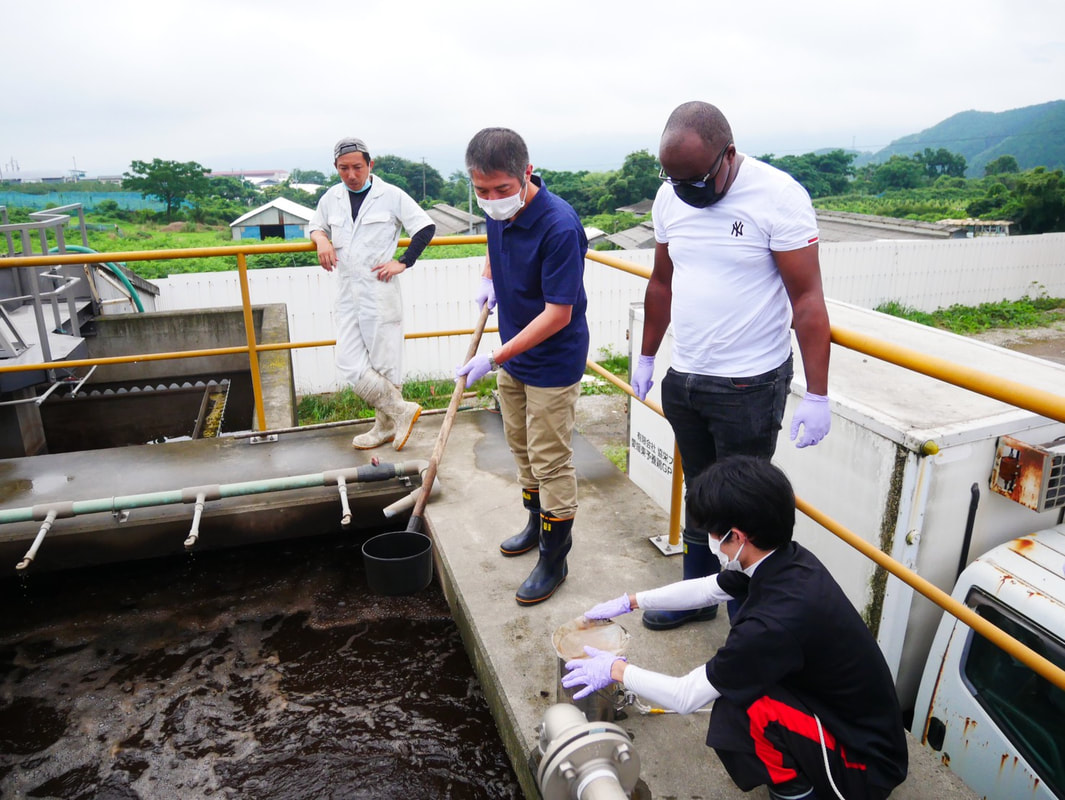
Sampling collection from wastewater treatment plant, river, and coastal area in Matsuyama, Ehime8/18/2022 The AMR team (Prof. Kozo Watanabe, Prof. Satoru Suzuki, Yuichiro Shimada, Ryota Ueda, Ngure Kagia, and Kenneth Bongulto) went for another sample collection from wastewater treatment plants, river, and coastal area in Matsuyama, Ehime. The sample collection aimed to collect water samples and isolate environmental bacteria and test for antimicrobial resistance genes (ARGs). Two different wastewater treatment plants were visited for the sampling. Water samples were collected from the wastewater influent and effluent. Environmental parameters were gathered per sampling site. Water samples were also collected from the coastal area and river where the wastewater effluent flows out.
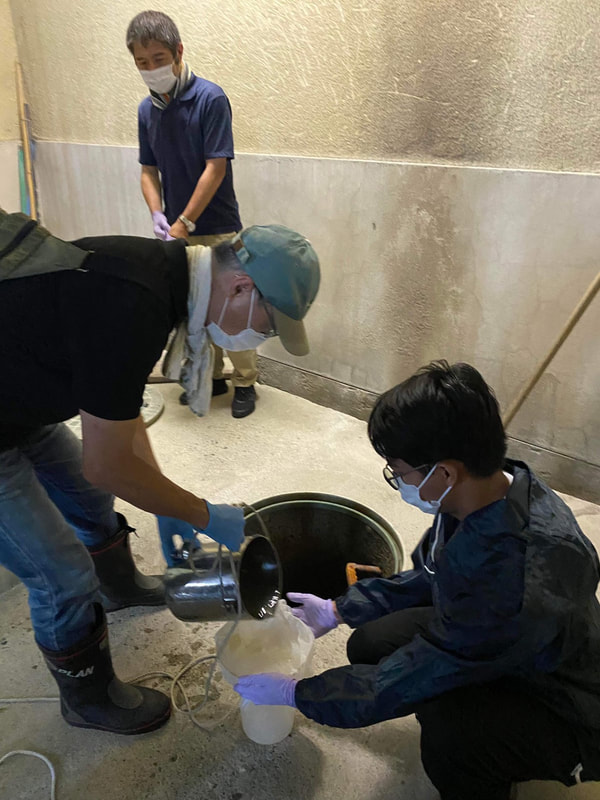
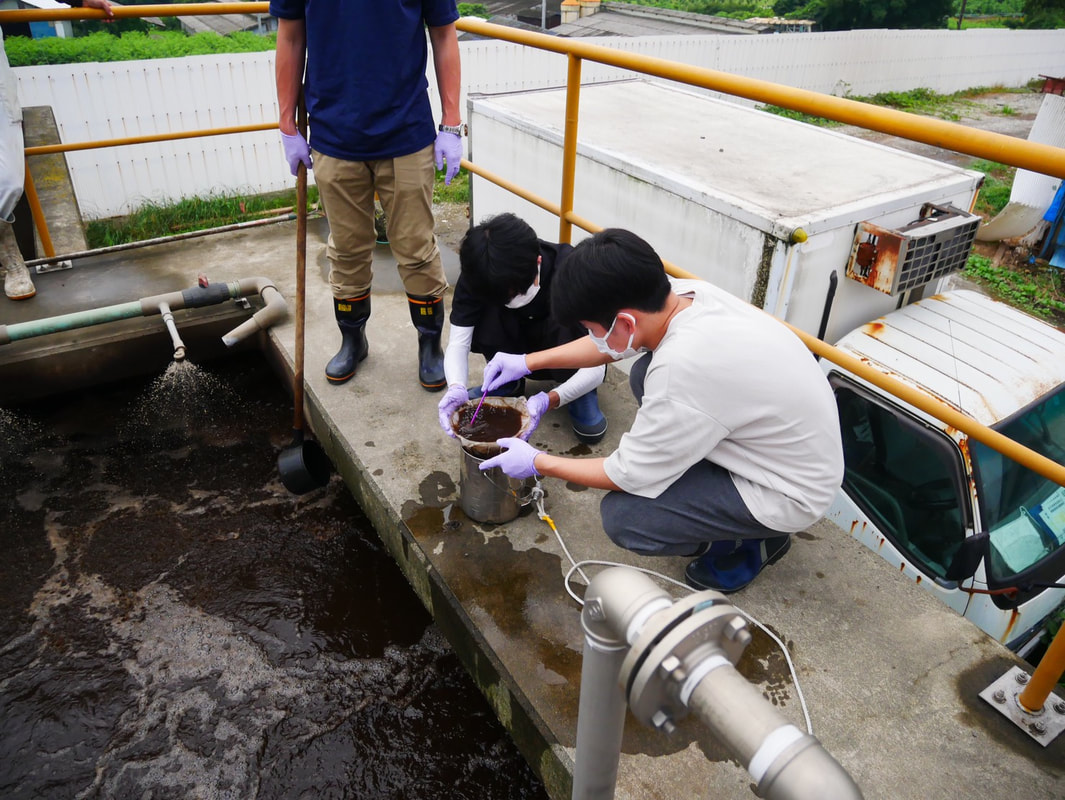
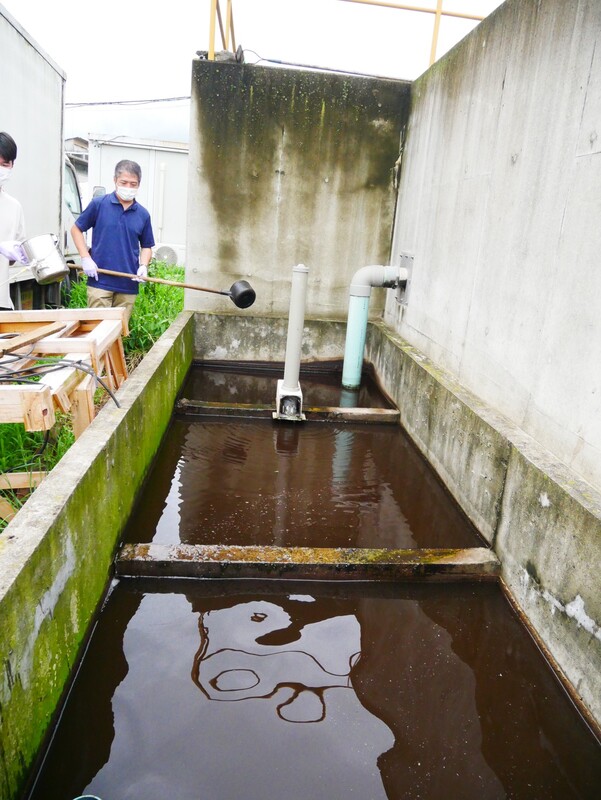
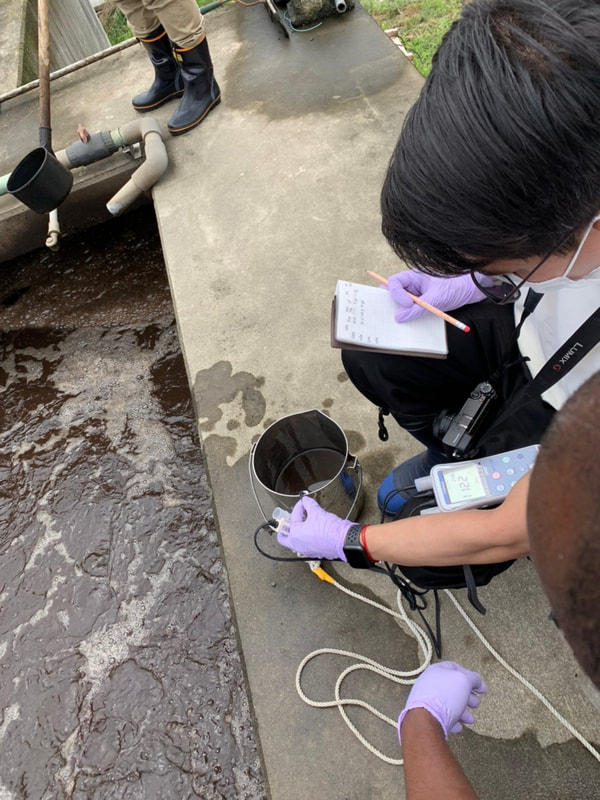
The AMR team (Prof. Kozo Watanabe, Yuichiro Shimada, Ryota Ueda, Ngure Kagia, and Kenneth Bongulto) collected wastewater samples from a pig farm last July 14, 2022. The aim of the sample collection is to isolate environmental bacteria and test for antimicrobial resistance genes (ARGs). Wastewater samples were collected from three different sections (influent, sequencing batch reactor, and effluent basin) of the wastewater treatment system. Environmental parameters were also gathered per sampling site.
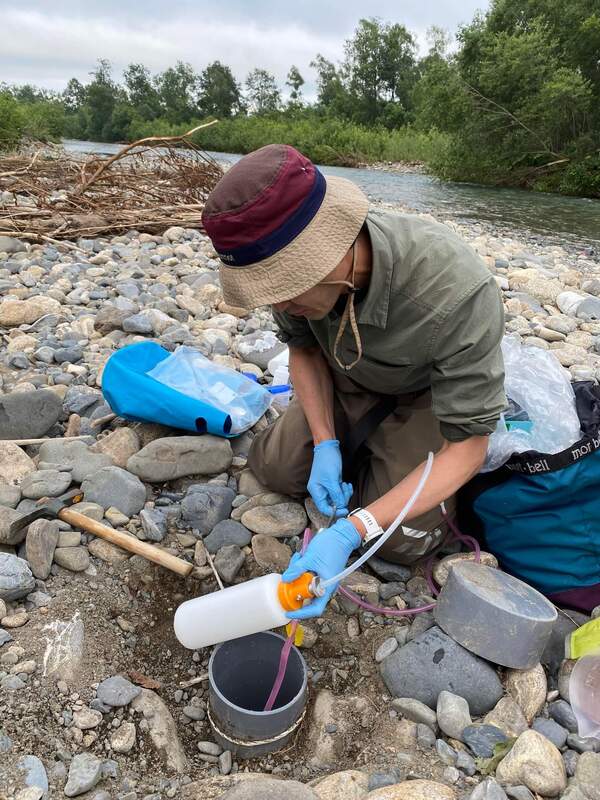
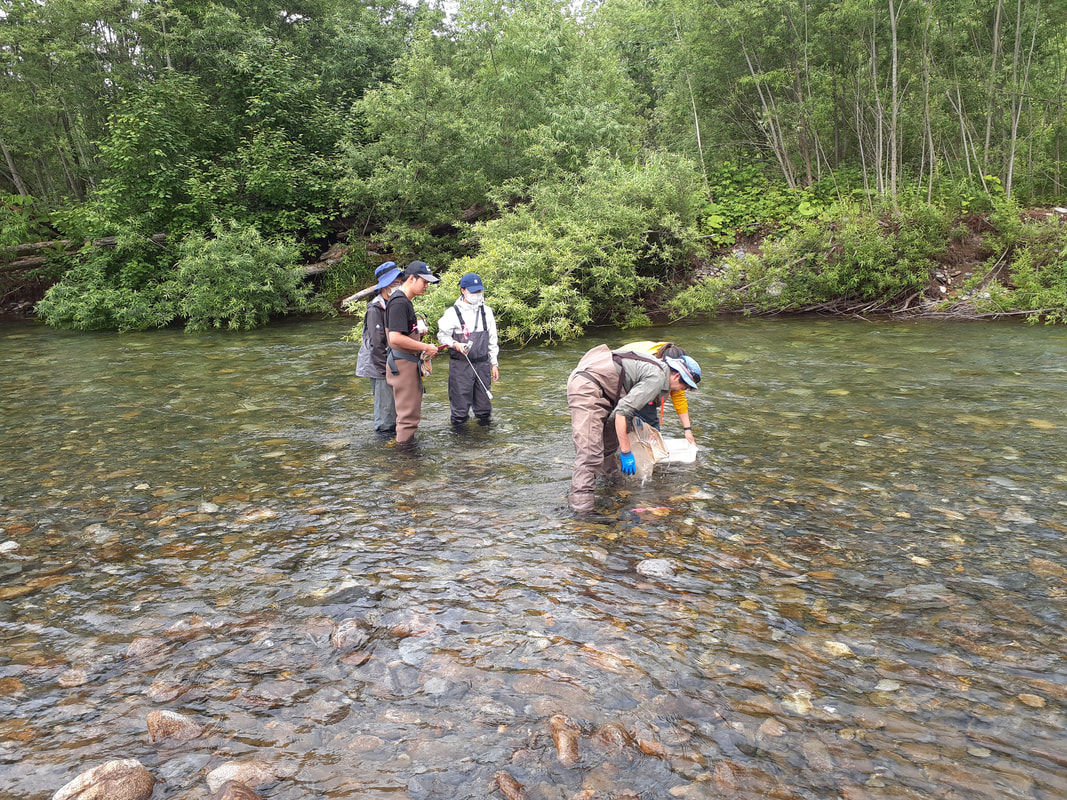
Environmental DNA (eDNA) sampling was performed again in Satsunai river, Hokkaido, Japan last July 6 to 10. Water samples for eDNA detection were collected from river and hyporheic zone. Macroinvertebrates both from benthic and hyporheic zones were also collected for morphological identification and DNA metabarcoding analysis. Professor Negishi from Hokkaido University performs water sampling from the hyporheic zone of Satsunai river for eDNA detection of macroinvertebrates. Water sample collected from the hyporheic zone were aseptically transferred to a sterile container for transport. Dan a Research student from MECOH lab performs water collection from the Satsunai river for eDNA detection. Dan together with the graduate students from Hokkaido University.
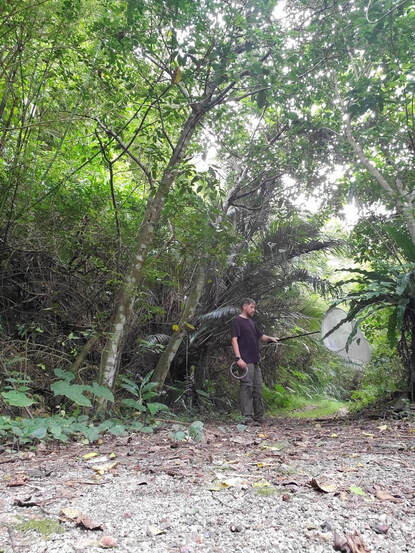
Dr. Levente-Péter Kolcsár (JSPS postdoctoral fellow) went on a field collection trip in the Japanese Alps, between 12-18 June. The aim of the trip was to collect crane flies (Diptera: Tipuloidea) along mountain rivers and streams for morphological and molecular studies. The main targeted group was a poorly known hairy-eyed crane fly genus (Tricyphona), in which several new species are suspected.
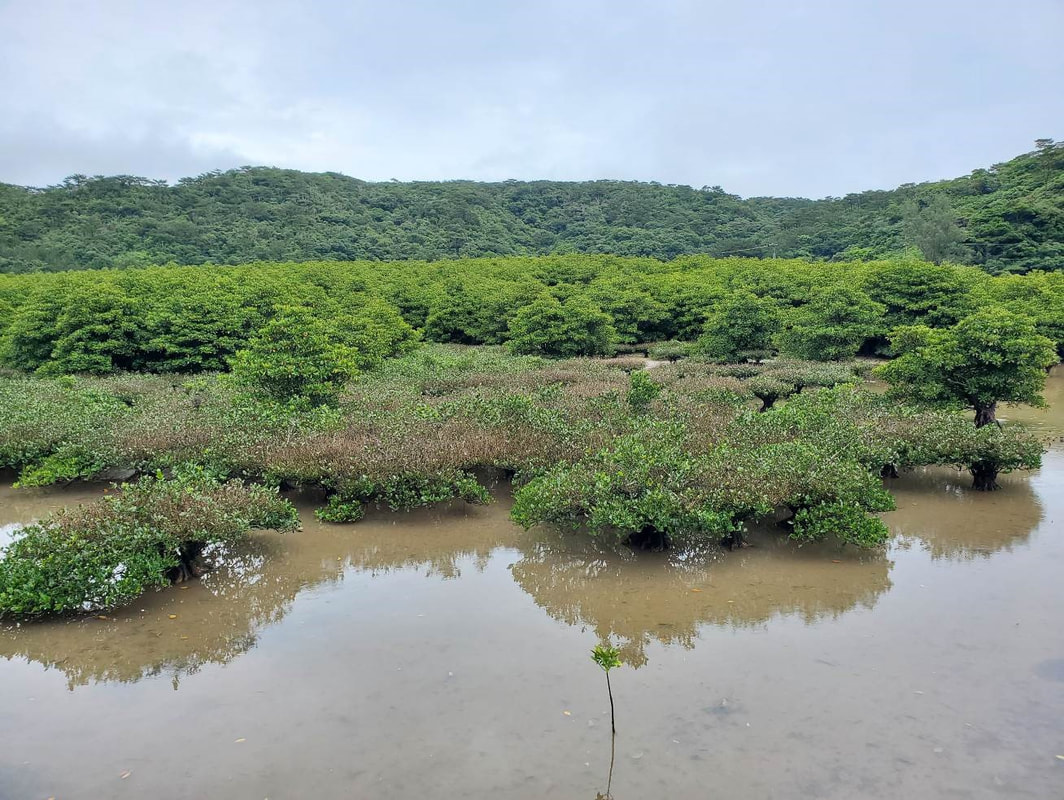
Dr. Daichi Kato from Echigo-Matsunoyama Museum of Natural Science, Matsunoyama, Niigata Pref. also joined the field collection between 14-16 June. During the trip over 1200 specimens belonging to 150-160 species were collected. Mr. Dan Joseph Logronio, a research student from the MECOH Lab performed eDNA collection in Satsunai river, Hokkaido, Japan last June 19 to assess macroinvertebrate communities both in benthic and hyporheic zones. DNA metabarcoding will be used for the identification of taxa present in the sample. This is a collaborative project with the Laboratory of Watershed Conservation and Management, Graduate School of Environmental Science, Hokkaido University headed by Dr. Junjiro Negishi.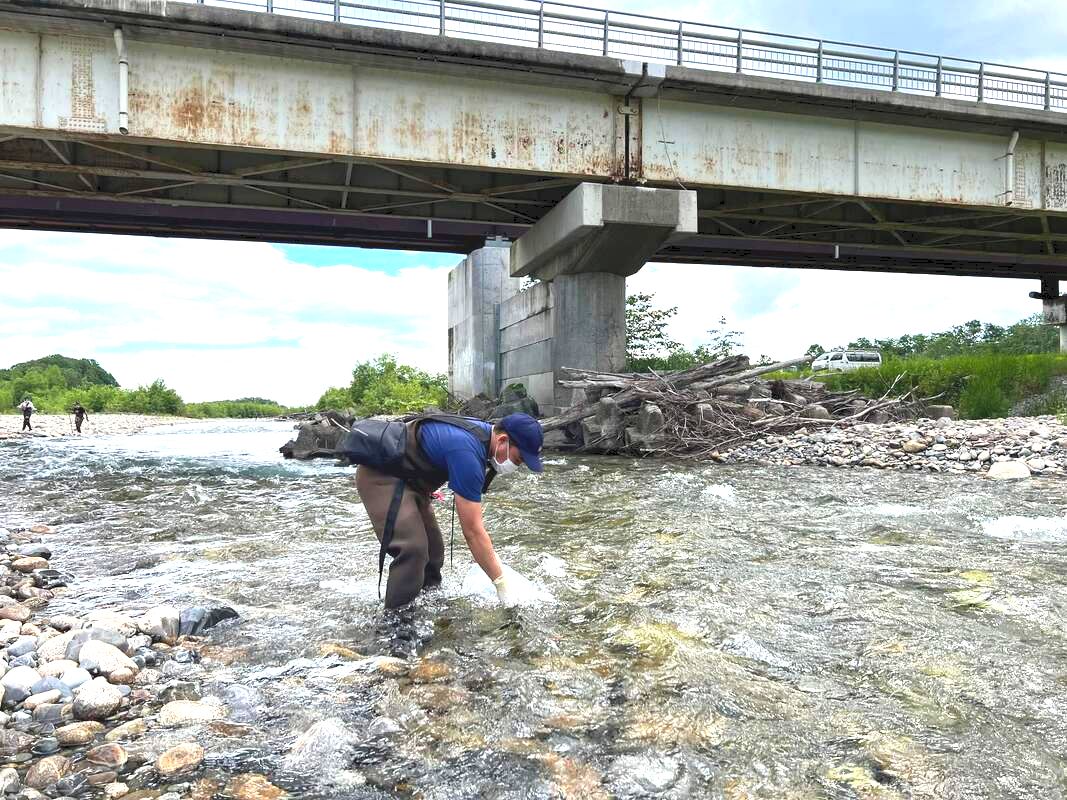  Dr. Levente-Péter Kolcsár (JSPS postdoctoral fellow) went on a field collection trip to Okinawa Island, Japan between 24-31 May, 2022. The aim of the trip was to collect crane flies (Diptera: Tipuloidea) along different aquatic and wet habitats. The crane fly fauna of the island is poorly investigated, and many species new to science or new to Japan are suspected from Okinawa and surrounding areas.
During the trip around 550-600 specimens belonging to 52 species were collected, from which 14 suspected to be new to science. Ngure Kagia from Kenya and Irish Coleen Asin from the Philippines joined the MECOH laboratory as new PhD candidates. Ngure finished his MSc. in Bioinformatics at the University of Leicester- Leicester UK. He has previously worked in the wet and dry laboratory on antimicrobial resistance (AMR) projects in Kilifi, Kenya and Cambridge, UK. Ngure has also worked on measles virus isolation in Nairobi, Kenya. His future interest in the MECOH laboratory revolve around the dynamics of antimicrobial resistant bacteria (ARB) in our environment. From the Philippines is Coleen who has obtained her Master of Science degree in Molecular Biology and Biotechnology at the University of the Philippines Diliman. She has worked on various research projects, one of which was surveillance of mosquito viruses thru mosquito viral metagenomics at UP Diliman. She has also worked on SARS-CoV-2 whole genome sequencing as a member of the Philippine Genome Center’s COVID-19 biosurveillance team. Going forward, she intends to focus and continue her research on mosquitoes and insect-specific viruses in the MECOH laboratory.
Khristina Judan Cruz (Karen) joins the MEcoH Lab as a postdoctoral fellow to do research work on the genome-wide signatures in genetically improved tilapia. She obtained her PhD in Biology at De La Salle University - Manila where she worked on the genetic characterization and gene expression analyses on Philippine genetically enhanced farmed tilapia. She plans to include comparative transcriptomics in her future projects. She is an Associate Professor at the Central Luzon State University, Science City of Munoz, Nueva Ecija, Philippines. Her research experience also includes integrating nanotechnology and bioactive compounds (nanobioapproach) for biomedical applications such as quorum sensing inhibition and anti-cancer action.
Kenneth and Dan are both new PhD candidates from the Philippines that recently joined the MECOH laboratory. Kenneth obtained his MSc in Biology at the De La Salle University - Manila, where he worked on nontuberculous mycobacteria isolated from patients with multidrug-resistant pulmonary tuberculosis (MDR TB). He has been part of research projects which revolved around tuberculosis, HIV, and dengue. For his future research projects, he plans to explore on the acquisition and mechanism of antimicrobial resistance genes (ARGs) in both clinical and environmental bacterial strains. Dan
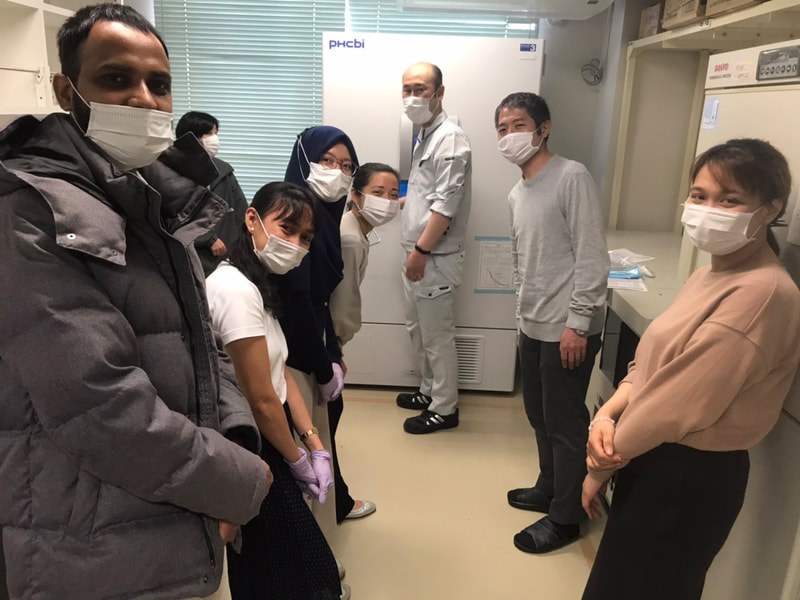
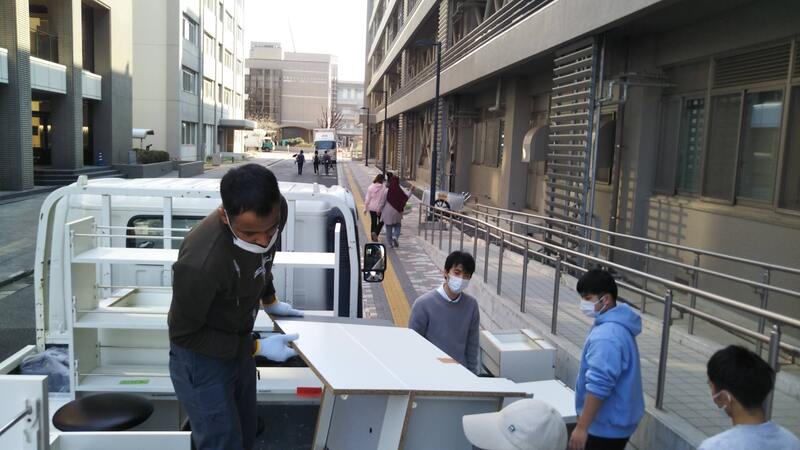
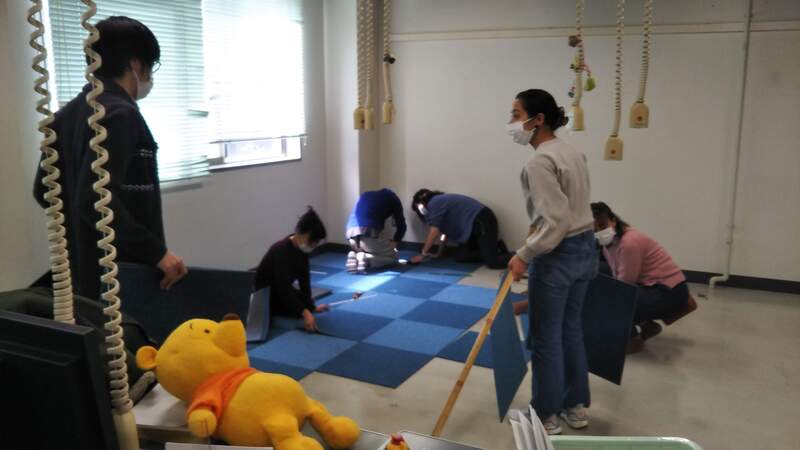
Dan finished his Master`s degree in Genetics minor in Molecular Biology and Biotechnology at the University of the Philippines - Los Baños (UPLB). He worked as Research Assistant and Associate Researcher at Southeast Asian Fisheries Development Center / Aquaculture Department (SEAFDEC/AQD) in the Philippines for 7 years. His research experiences include aquaculture, fish health and fish genetics. For his future research projects, he plans to explore the applications of DNA metabarcoding, population genomics and transcriptomics for the conservation and management of aquatic resources. Last March 7, the Molecular Ecology and Health Laboratory (MECOH) has moved to the General Research Bldg. 2, where the Division of Ecosystem Health Sciences of the Center for Marine Environmental Studies (CMES) is located. Although it was only a 150-meter away from its old home, we all worked hard together and managed to accomplish the mission. Now MECOH is part of CMES, both in name and in reality. The space is more than three times larger than before. It is very helpful because the number of people and lab equipment is increasing. MECOH family is growing! Here are some photos of the new MECOH laboratories and its members.
|
MECOH LabLatest news Archives
March 2024
Categories |























































































































 RSS Feed
RSS Feed